IGEM:Peking/2007/Count:Knockout
From OpenWetWare
Jump to navigationJump to search
General Strategy
- Use plasmid pKO3 to generate precise deletions in chromosome of Escherichia coli through homologous recombination
- For working principle of pKO3 please refer to Link A. et al., (1997) J Bacteriol, Vol. 179 (20), 6228–6237
- All pKO3 containing strains must be cultivated in the following condition to ensure the survival of plasmid pKO3: LB, Cm+ (25 μg/mL), 30℃ for 24 hours.
oriT Knock Out
oriT-deleted fragment construction
- oriT-F, -R751 and -pSC101 are fragments in length of several hundred (bp). To delete these, fragments contain up- and downstream sequences (500 bp each) but not oriT are constructed.
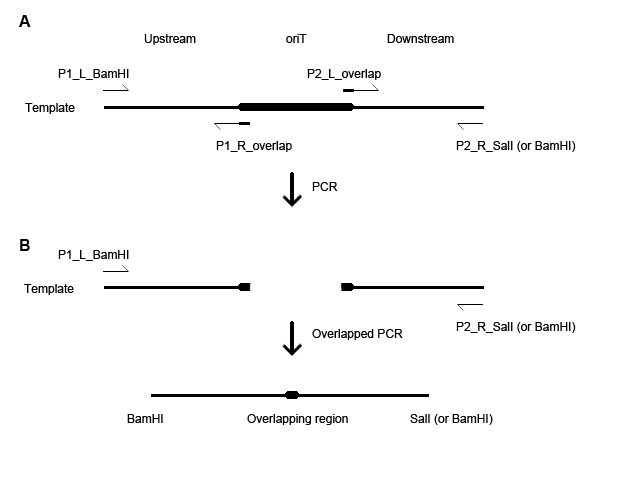
- For each oriT-deleted fragment construction, 2 pairs of primers are designed: each contains a 30-bp overlapping sequence (Fig. 1A).
- oriT-F has BamHI restriction sites at both of its 5’ and 3’ ends. oriT-R751 and oriT-pSC101 has BamHI at 5’ and SalI at 3’ end.
- Two sets of PCR are needed to produce a final product (Fig. 1B).
Sequencing fragments
- PCR fragments are digested by restriction enzymes. Hold the systems in 30℃ (F) or 37℃ (R751 and pSC101) for 20 hours.
- Ligate the digested fragments with pUC18 vector, then transformed them into DH5a competent cells and cultivate on Amp+ X-gal LB plates.
- Pick up white spots and send them to Boshang Zhanxin company for sequencing.
pKO3 plasmid construction
- Pick up sequencing-corrected clones and purify their plasmid pUC18.
- Digest using restriction enzymes and use gel separation and purification to acquire correct oriT-deleted fragments.
- Ligate the correct fragments with plasmid pKO3, then transformed them into DH5a competent cells and spread on Cm+ LB plates. Hold the plates in 30℃ for 24 hours.
- Pick up colonies for colony PCR verification with two primers: one from pKO3 (i.e. pKO3-R) and another from oriT-deleted fragment (i.e. oriT-F-1S). Make sure they cover different directions of the plasmid.
- Cultivate PCR-verified clones, purify their plasmid and digest using restriction enzymes for final verification.
Homologous Recombination
- Transform plasmid pKO3 with oriT-deleted fragments into competent cells with plasmid F, R751 and pSC101. Hold them in 30℃ for 24 hours.
- Pick up one colony and shake in liquid Cm+ LB for 12 hours.
- Spread the liquid onto 17μg/mL Cm+ plates, then hold in 43℃ for 12 hours. Set control in 30℃.
- Pick up one colony and shake in liquid Cm+ LB for 12 hours in 37℃.
- Spread the liquid onto Sucrose plates and hold in 30℃ for 24 hours for positive selection.
- Pick up colonies and draw lines in Cm+ LB plates for negative selection.
- PCR verification to get final products.