IGEM:UC Berkeley/2006/bryans notebook
Return to Bryan Hernandez
plate reader data order:
TG1, 23077, 129-3, 129-4, 124-2, 125-3, 123-2, 83-2, 82, 84, 83, 132-4, 132-2, 134-3, 81-4, 81-3, 81-2, 81-1
scanning key variant gel screening results:
remake ape files for new keys, ugh
Image1: gel1[ 98.1-.4, 99.1, 122.1-.4], gel2[ 123.1-.4, 124.1-.4], gel3[125.1-.4, 126.1-.4]
results: gel1[ 98.1-.4, 99.1, 122.1-.4]=all bad, gel2[123.2, 124.2]=look good, gel3[125.3]=looks good
Image2: gel1[127.1-.4, 128.1-.4], gel2[129.1-.4, 130.1-.4], gel3[131.1, 133.1-.4]
results: gel1[127.1-.4, 128.1-.4]=bad, gel2[129.3, 129.4]=look good, gel3=all bad
Image3: gel1[134.1-.4, 135.1-.4], gel2[132.1-.4, 136.1-.4]
results: gel1[134.3]=looks good, gel2[132.2, 132.4]=sequenceable
potentially successful clones:
sequenced:
- 83.2=good
- 81-1=bad, BB excision
- 81-2=good
- 81-3=good
- 81-4=good
- 123.2=bad, excision of BB
- 124.2=correct
- 125.3=bad, BB excision
- 129.3=sequencing failed. primer bound to 5' terminator
- 129.4=bad, BB excision
- 134.3=bad, triple insertion in sddressing sequence
- 132.2=bad, some sort of weird insertion
- 132.4=bad, very large insertion
probably failed clones
- 98
- 99
- 122
- 126
- 127
- 128
- 130
- 131
- 133
- 135
- 136
- (
Experiments
- got sequencing results back for 23081-84.
- 23081 lost some of its Ptet promoter...needs to be remade.
- 23082 good in biobricks
- 23083 looked weird. lost almost enitre biobrick. seq only showed the presence of b0015. looks like everything upstream of the actaga scar got removed. I sequenced the other two screens and they were correct. J23083-2 and J23083-3 are both correct.
- 23084 looked good inside biobricks
- transformants of 23098, 99, 122-136 came out kinda shitty. grew up about 4 colonies of each to screen. j23132 had no colonies so im retransforming it.
Experiments
--Bryanh 10:12, 1 December 2006 (EST)
- transformed pJ23006-J23098,99,122-136.
Experiments
--Bryanh 11:32, 29 November 2006 (EST) tRNA homology frame shifts
- repeated 5ml vs. 500ml experiment
- time course 3
note that 23081 and 23083 were not correct sequences
Box3
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| LB | LB | LB | TGI | TGI | TGI | 23077 | 23077 | 23077 | 23079 | 23079 | 23079 | |
| 23081 | 23081 | 23081 | 23082 | 23082 | 23082 | 23083 | 23083 | 23083 | 23084 | 23084 | 23084 |
Experiments
--Bryanh 16:12, 27 November 2006 (EST) tRNA homology frame shifts
- 5ml vs 500ml results: after 24 hours of growth and after 48 hours of growth
- prepped pJ23006-J23084-5. digest. insert.
- started second time course. see map below
box 2 time course run note that 23081 and 23083 were not correct sequences
Box2
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| LB | LB | LB | TGI | TGI | TGI | 23077 | 23077 | 23077 | 23079 | 23079 | 23079 | |
| 23081 | 23081 | 23081 | 23082 | 23082 | 23082 | 23083 | 23083 | 23083 | 23084 | 23084 | 23084 | |
| EZ | EZ | EZ | TGI | TGI | TGI | 23077 | 23077 | 23077 | 23079 | 23079 | 23079 | |
| 23081 | 23081 | 23081 | 23082 | 23082 | 23082 | 23083 | 23083 | 23083 | 23084 | 23084 | 23084 |
Experiments
--Bryanh 15:01, 26 November 2006 (EST) tRNA homology frame shifts
- make M9 supplemented media Endy:M9 media/supplemented
- grew up 23081, 82, 83, 84 + controls in EZ Rich & LB.
- ran time course. Note: the run stopped for a few hours because the wrong filter was installed. I swapped it with the correct filter and ran the protocol as is. This means that the cultures were sitting in the plate for ~3-4 hours at 37C.
- grew pJ23006-J23085-4 from plate. prep. digest (BamHI/HIndIII)
- extend new primers. digest BamHI/HindIII. insert
box1 time course run (.15uL into 150uL) note that 23081 and 23083 were not correct sequences
Box1
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| TGI | TGI | TGI | LB | LB | LB | 23077 | 23077 | 23077 | 23079 | 23079 | 23079 | |
| 23081 | 23081 | 23081 | 23082 | 23082 | 23082 | 23083 | 23083 | 23083 | 23084 | 23084 | 23084 | |
| TGI | TGI | TGI | 23077 | 23077 | 23077 | 23079 | 23079 | 23079 | EZ | EZ | EZ | |
| 23081 | 23081 | 23081 | 23082 | 23082 | 23082 | 23083 | 23083 | 23083 | 23084 | 23084 | 23084 |
Experiments
--Bryanh 22:51, 25 November 2006 (EST) tRNA homology frame shifts
- grow up 23081, 82, 83, 84 + controls in EZ Rich & LB.
- take OD.
- assay
- set up time course
- plated pJ23006-J23085-4 from -80. prep. digest (BamHI/HIndIII)
- extend new primers. digest BamHI/HindIII. insert
Experiments
--Bryanh 21:17, 22 November 2006 (EST) tRNA homology frame shifts--> J23082, J23083,
- Assay results
- (TGI, pSB3C6-J23077, pSB3C6-J23079-1): 343, 451, 85561,
- (pJ23006-J23081A): 644
- (pJ23006-J23082-1D): 6194
- (pJ23006-J23083-1D): 596
- (pJ23006-J23084-1D): 5761
- gain v distance from RBS
Experiments
--Bryanh 21:52, 17 November 2006 (EST)
tRNA.B0015-->J23081
- sequence
tRNA homology frame shifts--> J23082, J23083,
- Assay results
- (TGI, pSB3C6-J23077, pSB3C6-J23079-1): 364, 509, 85053
- (pJ23006-J23082-1A, B, C): 3616, 3798, 4125
- (pJ23006-J23083-1A, B, C): 571, 520, 556
- (pJ23006-J23084-1A, B, C): 7212, 6824, 6977
- gain v distance from RBS
orthogonal variants
- find a thermo model to predict orthogonality.
Experiments
--Bryanh 21:25, 15 November 2006 (EST)
J01022 RBS spacing fix --> J23079
- made a -80.
tRNA.B0015-->J23081
- sequence
tRNA homology frame shifts--> J23082, J23083,
- grew up 3 colonies of each cotransformant to assay
orthogonal variants
- find a thermo model to predict orthogonality.
Experiments
--Bryanh 21:35, 14 November 2006 (EST)
pSB3C6-J23079MUT
- grew up
- miniprep
- sequence entire biobrick
J01022 RBS spacing fix --> J23079
- transformed from miniprep,
- grew up colony
- Make a -80.
tRNA.B0015-->J23081
- sequence
tRNA homology frame shifts--> J23082, J23083,
- digest XhoI/AlwNI. successful clones will show one band
- cotransform with J23077 into TGI
- it appears that 23082-1,2; 23083-1; and 23084-1,2,3 were all good.
- send one of each for sequencing
orthogonal variants
- find a thermo model to predict orthogonality.
Experiments
--Bryanh 16:58, 12 November 2006 (EST)
J01022 RBS spacing fix --> J23079
- transform from miniprep, and make a -80.
tRNA.B0015-->J23081
- grew up the only colony to screen. fingers crossed.
tRNA homology frame shifts--> J23082, J23083,
- ligate in addressing sequence.
orthogonal variants
Experiments
--Bryanh 11:24, 26 October 2006 (EDT)
J01022 RBS spacing fix --> J23079
- sequencing came back good.
- sent chris pSb3C6-J23079-1
tRNA.B0015-->J23081
- grew up the only colony to screen. fingers crossed.
[tRNA precursor][B0015]-->J23085
- sequenced. came back perfect.
tRNA homology frame shifts--> J23082, J23083,
- awaiting 23085
Experiments
--Bryanh 06:06, 20 October 2006 (EDT)
J01022 RBS spacing fix --> J23079
- awaiting sequencing
tRNA.B0015-->J23081
- plasmid separation on pJ23006-J23070 x pSB3C6-J01122 TO GET J23070 out.
- pickd colony and miniprepd and digest SpeI/PstI.
- paste in b0015.
[tRNA precursor][B0015]-->J23085
- remapped colonies EcoRI/PstI.
tRNA homology frame shifts--> J23082, J23083,
- awaiting j23085
Experiments
--Bryanh 20:40, 6 October 2006 (EDT)
J01022 RBS spacing fix --> J23079
- transformed for the second time.
tRNA.B0015-->J23081
- plasmid separation on pJ23006-J23070 x pSB3C6-J01122 TO GET J23070 out.
- pickd colony and miniprepd and digest SpeI/PstI.
- paste in b0015.
[tRNA precursor][B0015]-->J23085
- mapped colonies EcoRI/PstI. they came out inconclusive because the gel resolution was so poor.
tRNA homology frame shifts--> J23082, J23083, J23084
- phusion extended oligos. digested BamHI/HindIII
Experiments
J01022 RBS spacing fix --> J23079
- saw no colonies.
- repeat subclone with gels. ran gel on J23079 XbaI/PstI digest. ligated. need to transform
- run gel on the pJ23076 SpeI/PstI digest.
tRNA.B0015-->J23081
- plasmid separation on pJ23006-J23070 x pSB3C6-J01122 TO GET J23070 out.
- pick colony and miniprep and digest SpeI/PstI.
- paste in b0015.
[tRNA precursor][B0015]-->J23085
- colony PCR: ca998/g00101
- positive: 620bp
- negative: 483bp
inconclusive. gel had extremely poor resolution. issues with EtBr
tRNA homology frame shifts--> J23082, J23083, J23084
- ordered oligos. b002-4
Experiments
--Bryanh 23:11, 30 September 2006 (EDT)
J01022 RBS spacing fix --> J23079
- miniprepped pJ23076 and digested to pasted in J23079.
- digested XbaI/PstI/DpnI and cleanuped J23079 PCR
tRNA.B0015-->J23081
- plasmid separation on pJ23006-J23070 x pSB3C6-J01122 TO GET J23070 out.
- miniprepped. diluted ~20 fold. transformed. plated on amp. plasmid is now isolated. miniprep and digest SpeI/PstI.
- paste in b0015.
[tRNA precursor][B0015]-->J23085
- miniprepped pJ23006-J23061. digested SpeI/PstI. ligated. transformed
tRNA homology frame shifts--> J23082, J23083, J23084
- ordered oligos. b002-4
Experiments
--Bryanh 22:31, 26 September 2006 (EDT)
- miniprepped the pSB1AK3-B0015
J01022 RBS spacing fix --> J23079
- grew up pJ23076 so that it can be miniprepped and digested to paste in J23079.
- digest and cleanup J23079 PCR
tRNA.B0015-->J23081
- cleaned up pSB1A2-b0015 PCR. digested and cleaned up.
- plasmid separation on pJ23006-J23070 x pSB3C6-J01122 TO GET J23070 out.
- grew up pJ23006-J23070 x pSB3C6-J01122
[tRNA precursor][B0015]-->J23085
- cleanuped pSB1A2-b0015 PCR. digested and cleaned up
- grew up pJ23006-J23061. digest SpeI/PstI. ligate. transform
tRNA homology frame shifts--> J23082, J23083, J23084
- order oligos. b002-4
Experiments
--Bryanh 19:22, 24 September 2006 (EDT)
J01022 RBS spacing fix --> J23079
- PCRd bh001 off of j01022.
tRNA.B0015-->J23081
- made construction file.
[tRNA precursor][B0015]-->J23085
- made construction file
tRNA homology frame shifts--> J23082, J23083, J23084
- made construction files.
JCAnderson 15:09, 6 September 2006 (EDT) Reads iG129-iG132
We got sequencing data for J23070 and J23062 (the ser2-derived keys). They are correct. I had to reconstruct some sequence files, though. The sequences for J23070, J23062, J23059, and J23060 in the registry are now all correct, as are the ApE files.
We also got sequencing of pSB3C6-J23071. Clone 1 was a failed read, but clone 2 was perfect. I don't know the status of pSB3C6-J23072 yet, but it is not yet sequenced.
JCAnderson 17:48, 24 August 2006 (EDT) I grew up colonies of pJ23006-J23066 (key3c/d.tt) with the lock3c and lock3d guys. I did this in 500 uL in 96 well blocks for a change. Not such a great idea, it comes out significantly differently than in well-aerated cultures:
<html xmlns:o="urn:schemas-microsoft-com:office:office" xmlns:w="urn:schemas-microsoft-com:office:word" xmlns:x="urn:schemas-microsoft-com:office:excel" xmlns="http://www.w3.org/TR/REC-html40">
<head> <meta http-equiv=Content-Type content="text/html; charset=windows-1252"> <meta name=ProgId content=Word.Document> <meta name=Generator content="Microsoft Word 10"> <meta name=Originator content="Microsoft Word 10"> <link rel=File-List href="1_files/filelist.xml"> <title>#1</title> <style> </style> </head>
<body lang=EN-US style='tab-interval:.5in'>
|
<o:p> </o:p> |
#1<o:p></o:p> |
#2<o:p></o:p> |
#3<o:p></o:p> |
<o:p> </o:p> |
value<o:p></o:p> |
subt<o:p></o:p> |
xfold<o:p></o:p> |
|
J23066/J01122<o:p></o:p> |
181<o:p></o:p> |
141<o:p></o:p> |
<o:p></o:p> |
<o:p></o:p> |
161<o:p></o:p> |
118<o:p></o:p> |
20<o:p></o:p> |
|
J23066/J23048<o:p></o:p> |
223<o:p></o:p> |
219<o:p></o:p> |
205<o:p></o:p> |
<o:p></o:p> |
216<o:p></o:p> |
173<o:p></o:p> |
29<o:p></o:p> |
|
J23066/J23049<o:p></o:p> |
346<o:p></o:p> |
<o:p></o:p> |
283<o:p></o:p> |
<o:p></o:p> |
315<o:p></o:p> |
272<o:p></o:p> |
45<o:p></o:p> |
|
<o:p> </o:p> |
<o:p></o:p> |
<o:p></o:p> |
<o:p></o:p> |
<o:p></o:p> |
<o:p></o:p> |
<o:p></o:p> |
<o:p></o:p> |
|
1129/1122<o:p></o:p> |
66<o:p></o:p> |
<o:p></o:p> |
<o:p></o:p> |
<o:p></o:p> |
66<o:p></o:p> |
23<o:p></o:p> |
4<o:p></o:p> |
|
1122<o:p></o:p> |
49<o:p></o:p> |
<o:p></o:p> |
<o:p></o:p> |
<o:p></o:p> |
49<o:p></o:p> |
6<o:p></o:p> |
1<o:p></o:p> |
|
1022<o:p></o:p> |
5954<o:p></o:p> |
<o:p></o:p> |
<o:p></o:p> |
<o:p></o:p> |
5954<o:p></o:p> |
5911<o:p></o:p> |
985<o:p></o:p> |
|
TG1<o:p></o:p> |
43<o:p></o:p> |
<o:p></o:p> |
<o:p></o:p> |
<o:p></o:p> |
43<o:p></o:p> |
0<o:p></o:p> |
0<o:p></o:p> |
<o:p> </o:p>
</body>
</html>
JCAnderson 12:20, 23 August 2006 (EDT)
Bryan had noticed that the rbs spacing in lock3 is not the same as b0034. We are therefore starting with (probably) a weaker rbs. I ordered oligos to make the proper-spacing variants of lock3 and lock3d in making parts J23071 and J23072 according to the construct file from 082006.
Also, I noticed that there is a hairpin at the 5' end of the new lock3D due to the XbaI site. I'm taking that out and also extending the 5' end with construct J23073, a variant of J23072.
JCAnderson 22:29, 21 August 2006 (EDT)
I assayed the J23062 and J23070's transformed into 1122:
- TG1 31
- 1022 3725
- 1122 41
- 1122/1129 78
- J23062-1A 121
- J23062-1B 121
- J23062-1C 128
- J23062-2A 151
- J23062-2B 140
- J23062-2C 113
- J23070-1A 297
- J23070-1B 317
- J23070-1C 309
- J23070-2A 410
- J23070-2B 39
- J23070-2C 44
So, it looks like J23070 clone 1 is solid, slightly more active than was key3d. Will sequence clone 1 of both j23062 and j23070 to confirm.
JCA081906--Half the group left friday, nobody's around for the next week, so I cleaned up the current stuff: The sequencing for pJ23006-J23061 (the tRNA BamHI-XhoI-HindIII guy) came back good (reads iG126 and iG127). Clone 1 is a perfect match to the model, clone 2 duplicates the XhoI/HindIII fragment, but inserting things into BamHI/HindIII would still give the desired products.
Bryan and Samantha left 4 cultures each of the tRNA-like keys J23062 and J23070. I did colony PCRs with CA998 and G00101 on the cultures, then digested each with XhoI. No cutting is a good clone. The gel goes:
- J23062 without XhoI
- Marker
- (Rest have XhoI)
- J23062 #1-->#4
- J23070 #1-->#4
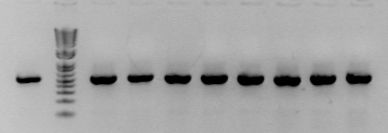 So, all look good. I'm miniprepping clone 1 and 2 of each, transforming 1122 to assay.
So, all look good. I'm miniprepping clone 1 and 2 of each, transforming 1122 to assay.
2.
- pick colonies of pJ23019-J23031-1 x pSB1A2-J01129 and pJ23019-J23032-1 x pSB1A2-J01129 to assay tomorrow.
- colony PCR pSB3CG0-E1010 and then grow up the good colonies so that they can be miniprepped tomorrow.
3.
- analyze sequence data.
8.
- pick like 4 more colonies from the original transform plate to grow up and miniprep again. start the subclone over.
9.
- grow up controls for assay plus regrow pJ23006-J23066-1 x pSB3C6-J01122
10.
- digest and map pJ23006-J23061-1,2,3,4 with AlwNI/XhoI
- send clones 1&2 for sequencing
- digest the correct clone with BamHI/HindIII
- ligate with the J23059 and J23060
- transform.
1.pJ23006-J23009-1 assay
2.Lock3 Variants
3.pK2, pK3--1129 assays
4.QPCR Experiment
5.key3c-key3d juxtaposation
6.Key3d loop adjustments
7.key3d-TT construct
8.pJ23006 GFP insert fix for registry
9.[key3c][key3d][B0015] construction
10.tRNA key
2.
- analyzed sequence data for pSB3C6-J23031-1, pSB3C6-J23032-1, and pSB3C6-J23036-1. pSB3C6-J23036-1 turned out to be very wrong. Apparently there was some weird genomic DNA amplification that somehow ligated itself into the plasmid. construction will start again.
- transformed pJ23019(pSB3C6)-J23031-1 and pJ23019-J23032-1 into 1129.
- threw away original clone plates of pSB3C6-J23030-3, pSB3C6-J23033-4, pSB3C6-J23035-A (here pSB3C6=pJ23019)because they appear to all be parent vector. construction of these will be restarted but instead with a new strategy. I dropped a genimiacin (sp?) cassette into the EcoRI/XbaI of pJ23019 so that the screening steps will be easier and the larger band cut out will be more evident.
3.
- analyze sequence data
6.
- picked colonies to grow up and assay along with the proper controls.
8.
- grow up pSB1A2-I13504 to miniprep
9.
- miniprep pJ23006-J23066-1 and send for sequencing.
- transformed into 1122.
10.
- PCR extended the KB032/KB033 oligos.
- did not map
- cleanup. digested.
- cleanup. ligated into pJ23006 XbaI/PstI.
- transformed. and then screen with XhoI/AlwNI tomorrow.
--Bryanh 17:12, 14 August 2006 (EDT)
1.pJ23006-J23009-1 assay
2.Lock3 Variants
3.pK2, pK3--1129 assays
4.QPCR Experiment
5.key3c-key3d juxtaposation
6.Key3d loop adjustments
7.key3d-TT construct
8.pJ23006 GFP insert fix for registry
9.[key3c][key3d][B0015] construction
7.
- PCR screen to verify the presence of B0015 came out positive. sent pJ23006-J23022-1 for sequencing using the reverse sequencing oligo BBa_G00101 to verify the junction between the J23009 and B0015 parts.
9.
- a PCR screen to confirm the presence of B0015 came out positive. will send pJ23006-J23066-1 for sequencing using the reverse sequencing oligo BBa_G00101 to verify the junction between the J23043 and B0015 parts.
- grow up 23066-1
- made a shitload 1129 competent cells.
- grew up pSB3C6-J23030-3, pSB3C6-J23033-4, pSB3C6-J23035-A (here pSB3C6=pJ23019)
- run gel for the pJ23006-J23059 and pJ23006-J23060 screen. transform the a correct one.
- transormed pSB1A2-I13504 instead of pSB1A2-I13500 because it wasnt working.
- colony PCRd the pJ23006-J23066 original clones. all of them came out looking perfect. grew up pJ23006-J23066-1 to miniprep, sequence and transform into 1122.
--Bryanh 22:01, 11 August 2006 (EDT)
1.pJ23006-J23009-1 assay
2.Lock3 Variants
3.pK2, pK3--1129 assays
4.QPCR Experiment
5.key3c-key3d juxtaposation
6.Key3d loop adjustments
7.key3d-TT construct
8.pJ23006 GFP insert fix for registry
9.[key3c][key3d][B0015] construction
- start the 1129 competent cell production.
- continue lock3variant production by sequencing pSB3C6-J23031-1, pSB3C6-J23032-1, and pSB3C6-J23036-1.
- digest pJ23006-J23030-3, pJ23006-J23033-4, pJ23006-J23035-A EcoRI/SpeI and ligate with the pSB3C6-J23018 EcoRI/XbaI digest.
- miniprep and screen pJ23006-J23059 and pJ23006-J23060 using a Ptet forward oligo and G00101. transform the ones that look good into 1122 and dont bother sequencing because its not going to work anyway.
- assay pJ23006-J23022.
- grow up pSB1A2-I13500
- Digest pSB1AK3 (XbaI/AlwNI, Neb2, small-761)(im pretty sure this already exists) Paste into pJ23006-J23043 (XbaI/AlwNI, Neb2, large-1724)(however this one doesnt.) and ligate them.
2.
- grew a 10ml cultures of pSB1A2-J01129
- threw out all pJ23006-lock3variants that are bad after analyzing XbaI/AlwNI digest. 23034 is the only one that we dont have at least one correct clone. forget about it for now unless the others come out particularly interesting.
3. tecan results for pSB1A2-J23041-1 and pSB1A2-J23042-1:
TGI: 23, 1122: 37, 1122x1129: 53, 1022: 3634
41A: 96, 41B: 96, 41C: 86
42A: 125, 42B: 141, 42C: 141
4.
- we are waiting for a kit to do this with.
5. tecan results for pJ23006-J23043-1.1 and pJ23006-J23044-1.1:
TGI: 23, 1122: 37, 1122x1129: 53, 1022: 3634
43A: 286, 43B: 272, 43C: 284
44A: 217, 44B: 185, 44C: 66
6.
- grew up 4 white colonies of each pJ23006-J23059 and pJ23006-J23060 to miniprep, screen, and transform into 1122.
7.
- grew up 1 colony of pJ23006-J23022-1 to tecan (along with all controls)
8.
- transformed pSB1A2-I13500 from DNA-1 box into TGI to grow up.
9.
- made construction file: pJ23006-J23066
1.pJ23006-J23009-1 assay
2.Lock3 Variants
3.pK2, pK3--1129 assays
4.QPCR Experiment
5.key3c-key3d juxtaposation
6.Key3d loop adjustments
7.key3d-TT construct
2.
- miniprepped pJ23006-J23033-D, pJ23006-J23033-E, pJ23006-J23033-F, pJ23006-J23034-D, pJ23006-J23034-E, pJ23006-J23034-F. test digest these with XbaI/AlwNI.
- 23030, 23031, 23032, and 23036 need to be tecanned agaisnt 1122 and need to be mapped.
- 1129 was replated and will be grown up for competent cell production.
3.
- sequenced pSB1A2-J23041-1 and pSB1A2-J23042-1
- picked three colonies of each pSB1A2-J23041,42 x 1122 to grow up and tecan.
4. talk to john
5.
- tecan the pJ23006-J23043-1 x J01122 (the first time this was transformed the 44 version didnt work so i completely redid both which is why im assaying 43 twice and 44 only once.)
- picked three colonies of each pJ23006-J23043,44 x 1122 to grow up and tecan again.
6.
- digested J23059 and J23060 with BamHI and HindIII
- miniprepped pJ23006-J23058-1 and purified the BamHI/HindIII digest.
- ligated to make te 23059 and 23060 constructs.
- transformed
7.
- transformed pJ23006-J23022-1 into 1122s.
--Bryanh 03:54, 9 August 2006 (EDT)
1.pJ23006-J23009-1 assay
2.Lock3 Variants
3.pK2, pK3--1129 assays
4.QPCR Experiment
5.key3c-key3d juxtaposation
6.Key3d loop adjustments
7.key3d-TT construct
1.
- discuss sequence results with chris
2.
- update map for pSB3C6-J23018 according to new sequence data.
- miniprepped pSB3C6-J23031-1, pSB3C6-J23032-1, pSB3C6-J23036-1
- picked colonies for pSB3C6-J23031-1, pSB3C6-J23032-1, pSB3C6-J23036-1 along with J01122 to compare.
- investigate the locks that didnt clone correctly.
3.
- colony PCR off cultures.
For pSB1A2-J23041 and pSB1A2-J23042: ca998/G00101
clone: 343bp parent(pSB1A2-J01007):238bp parent(pJ23006-J23008):474bp
all cultures were perfect. miniprep pSB1A2-J23041-1 and pSB1A2-J23042-1
- transformed into 1122
4.
- fix KB oligo/QPCR oligo naming overlap.
5.
- resequenced pJ23006-J23044-1 using this time G00101
- picked a colony to grow and tecan
6.
- grew up one green colony.
7.
- colony PCR a few colonies of this using ca998/G00101 as the primers and pJ23006-J23009 as the parent control.
clone: 692bp parent(pJ23006-J23009): 345bp parent(pSB1AK3-B0015): 388bp
all colonies were perfect.
- grew up colony of pJ23006-J23022-1.
Bryanh: 08/08/06
- pJ23006-J23009-1 assay
- Lock3 Variants
- pK2, pK3--1129 assays
- QPCR Experiment
- key3c-key3d juxtaposation
- Key3d loop adjustments
- key3d-TT construct
- jen sequenced pSB3C6-bca1025 with ca1001 and filled out the sequence log as "kad"
1.
enter sequencing results.
2.
- miniprep pSB3C6-J23031-1, pSB3C6-J23032-1, pSB3C6-J23036-1;
- miniprepped pJ23006-J23030-B,C, pJ23006-J23033-A,B,C, pJ23006-J23034-A,B,C, pJ23006-J23035-A,B,C. test digested with XbaI/AlwNI. for a successful sublcone i expect to see bands of ~600, ~1700bp. a singley cut plasmid results in a band of ~2300bp. one band at about 1150bp is doubley cut where both bands are on top of each other.
- transformed pSB3C6-J23031-1, pSB3C6-J23032-1, pSB3C6-J23036-1
3.
picked colonies of pSB1A2-J23041 and pSB1A2-J23042
4.
hold off
5.
- miniprepped and sequenced pJ23006-J23043.1 and pJ23006-J23044.1
- transformed into 1122
6.
- miniprepped pJ23006-J23009-1. Digested pG80-GFPmut3 (BamHI/HindIII, Neb2, small-887bp),
Pasted into pJ23006-J23009-1 (BamHI/HindIII, Neb2, large-2323bp).
- transformed
7. digested, ligated, and transformed key3d with b0015 according to the construct file pJ23006-J23022.
--Bryanh 18:46, 7 August 2006 (EDT)
- pJ23006-J23009-1 assay
- Lock3 Variants
- pK2, pK3--1129 assays
- QPCR Experiment
- key3c-key3d juxtaposation
1. analyze sequence data
2.
- grew up clones that were correct according to the sequencing. pJ23006-J23031-1, pJ23006-J23032-1, pJ23006-J23036-1 were correct.
- picked 3 new colonies for pJ23006-J23030-1, pJ23006-J23033-1, pJ23006-J23034-3, pJ23006-J23035-1. these will be miniprepped and then mapped with XbaI/AlwNI. if bad then we expect to only see one fragment because XbaI is deleted, among other things.
3. miniprepped and digested pJ23006-J23008 and then ligated with the J01007/J01006 digests. transformed.
4.
hold off.
5.
Colony PCR
23043.1-43.4 and 23044.1-44.4 is ca998 and g00101. one control using same oligos but 23006-23008 as a template. all of these came out looking the right size.
- pick a colony from the first of each colony.
8-4-06
hey brian.
we ligated and transformed J23043 and J23044 for you today. we couldn't make J23021 and J23042 though with the J01006 and J01007 digests because there wasn't a J23008 Alwn/Xbal digest, only a J23008 Alwn/Spe digest and there was not enough of the J23008 miniprep to make a new digest, so we grew up J23008 from the -80 instead. Also, you guys are running low on J23009 but didn't have a minus 80 stock of it, so me and matt transformed the miniprep that was sequenced in TGIs.
-Samantha
Hey Brian. This is what I did for you today:
- religated key3 variants with pSB3C6-J23018 (it wouldnt have worked to plate your ligation/transformations becuase they sat in the shaker over night)
- transformed and plated on CAM plates
- made names for all 7 of these constructs and temporary construction files (need to be formatted correctly. i just put into in there so we know what they are)
- checked sequencing results, threw out transformations i did earlier of pJ23006-J23035-1, pJ23006-J23034-3, pJ23006-J23033-1, and pJ23006-J23030-1. picked 3 new colonies of each of those.
--Bryanh 20:54, 3 August 2006 (EDT)
- pJ23006-J23009-1 assay
- Lock3 Variants
- pK2, pK3--1129 assays
- QPCR Experiment
- key3c-key3d juxtaposation
1.
- tecan assay:
199 287 431 22903 1388 1309 867 973 1214
TGI, 1122, 1122x1129, 1022, pJ23006-J23009-1-E, pJ23006-J23009-1-F, pJ23006-J23009-1-G, pJ23006-J23009-1-H, pJ23006-J23009-1-I
- sent for sequencing again using the reverse sequencing oligo G00101.
2.
Digested the lock3variant PCRs and ligated with the 23018 digest.
- transformed but not plated. the cells are in the rescue phase incubating in LB. these just need to be plated on chloramphenicol now.
- sent pSB3C6-J23018 for sequencing off the midiprep.
3.
- fixed construction error. experiment needs to be restarted.
4.
- hold off for now.
5.
- digested pJ23006-J23008 and pJ23006-J23009 according to the construct files for pJ23006-J23043 and pJ23006-J23044.
i gel purified these digests so that they are ready to be ligated as is. this means that the Digest pJ23006-J23008 (AlwNI/SpeI, Neb2, large-1826bp) and Digest pJ23006-J23009 (AlwNI/XbaI, Neb2, small-652bp are cleaned up and mixed together and ready to be ligated to make pJ23006-J23043. likewise the pJ23006-J23044 construct is in the same form. these ependorfs are labeled as "pJ23006-J23043 dig. C" and "pJ23006-J23044 dig. C" and are sitting in the riboregulator's oligonucleotide box (since this is the only one with room.)
--Bryanh 23:54, 1 August 2006 (EDT)
experiments
- pJ23006-J23009-1 assay
- Lock3 Variants
- pK2, pK3--1129 assays
- QPCR Experiment
1.assayed: <> 6 7 8 9 10
A 257 444 729 2287 56416
TGI, 1122, 1122x1129, pJ23006-J23009-1x1122, 1022
These results are somewhat unexpected compared to yesterday's assay which was pretty much the same thing except we have 1022 data on this one.
- due to the variation in flouresence the experiment will be repeated using several different colonies in an effort to establish relatively presice numbers.
2. minipreped pSB3C6-J230181,2. digested EcoRI/XbaI. the gel came out really bad due to the low copy number of the p15a backbone and the digest of the 23006-lock3variants also came out really bad because the fragment was so small it wasnt visible on the gel (~260). as such, a pcr strategy will be taken to amplify the small fragment using the pSB1A2 sequencing oligos (ca998 and G00101) and then digest from there. the pcr has been done. in order to get the pSB3C6-J23019-1 backbone digested we will need to midiprep it to obtain a high concentration of the low copy plasmid and then the digest should come out better.
- a 50ml solution of pSB3C6-J23018 has been grown up for the midi.
- sent one of each 23006-lock3variant for sequenceing.
3. grew up two colonies of each pSB1A2-J230024,-J23025
--Bryanh 31 July 2006 (EDT)
experiments
- pJ23006-J23009-1 assay
- Lock3 Variants
- pK2, pK3--1129 assays
- QPCR Experiment
1. assay: 210, 449, 248, 312 (TGI, 1122, 1122x1129, pJ23006-J23009-1)
2. grow up pSB3C6-J23018. screen pJ23006-Lock3Variants. miniprep them.
PCR Screen:
pJ23006-J23030: KB014/Bba_G0010 expected band size: ~140 pJ23006-J23031: KB014/Bba_G0010 expected band size: ~140 pJ23006-J23032: KB014/Bba_G0010 expected band size: ~140 pJ23006-J23033: KB018/Bba_G0010 expected band size: ~140 pJ23006-J23034: KB020/Bba_G0010 expected band size: ~140 pJ23006-J23035: KB021/Bba_G0010 expected band size: ~140 pJ23006-J23036: KB022/Bba_G0010 expected band size: ~140
3. digested J01007, J01006 with AlwnI/SpeI and ligated with pSB1A2-J01129.transformed.
4. oligos arrived
for monday:
1.grow up colonies.
2. colony PCR pJ23006-lock3Variants and then miniprep the good ones. digest EcoRI/SpeI (this doesnt have to be done on this day.)
3. miniprep J01007, J01006. digest and ligate into pSB1A2-J01129. tranform.
--Bryanh 15:22, 27 July 2006 (EDT)
experiments
- pJ23006-J23009-1 assay
- Lock3 Variants
- pK2, pK3--1129 assays
- QPCR Experiment
1. TGI, 1122, 1122x1129, pJ23006-J23009-1 plates are sitting in the fridge.
2. grew up four colonies of each pJ23006-Lock3Variant to colony PCR (or miniprep PCR) them on monday.
3. minipreped J01007, J01006. Digestion coctail is ready to go and is sitting in the fridge, just add enzymes (AlwnI and SpeI when its here.)
4. waiting on oligos
--Bryanh 15:24, 26 July 2006 (EDT)
experiments
- pJ23006-J23009-1 assay
- Lock3 Variants
- pK2, pK3--1129 assays
- QPCR Experiment
1.
- pJ23006-J23009-1 x J01122 is sitting on a plate.
- transformed 1122x1129 and plated TGI and pSB3C6-J01122.
2.
- grew two colonies of pSB1A2-J23018
- Klenow extension of lock3 variants:
- J23030: KB014/KB015
- J23031: KB014/KB016
- J23032: KB014/KB017
- J23033: KB018/KB019
- J23034: KB020/KB019
- J23035: KB021/KB019
- J23036: KB022/KB019
3. J01007, J01006 cultures are growing up.
4. ordered oligos for QPCR --Bryanh 15:22, 27 July 2006 (EDT)