OhioMod2013:Introduction
Background
DNA origami is a relatively recent technique first described in 2003 as a method of folding DNA into complex structures [1]. Since then there has been an effort to find practical applications for DNA origami, many of which involve targeted in vivo delivery of drug payloads [2], antigens [3], and siRNA [4]. We believe however, that the strongest potential is for delivery of the DNA itself by incorporating the gene vector into the origami for cellular uptake.
The viral method is the currently the most efficient at transfecting mammalian cells due to the intrinsic ability of the viral vector. However because of the immunogenic properties of the viral vehicle and its high production costs, artificial non-viral methods are being investigated. Inorganic nanoparticles are especially tantalizing due to their synthetic production, durable storage, and resistance to premature degradation or digestion. The most common types of nanoparticles include cationic polymers, liposomes, gold clusters, and silicate particles. Among them, two successful commercial agents are Polyfect® (dendritic polymer) and Lipofectamine® (liposome). To have transfection efficiencies comparable to these two products would be considered our standard of success.
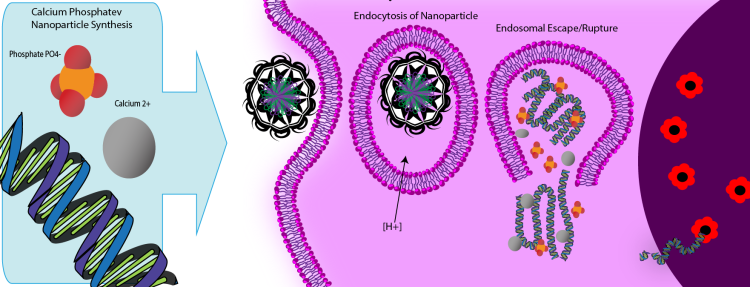
One promising material that has yet to be fully realized is that of the calcium phosphate nanoparticle (CPNP). First described 35 years ago[5], Calcium phosphate is non-toxic, easily biodegradable, and is simply precipitated in aqueous solutions. The greatest technical challenge in their production is ensuring stable uniform sizes, as they can easily agglomerate without proper dispersion. Many labs have gone on to show that CPNP improves the uptake of their active agents into the cell, whether they be flourophores [6], antigens and immunoactive oligonucleotide [7], or therapeutic genes [8]. This improved uptake by CPNPs can be due to a number of reasons, including the size of the nanoparticle, the agglomeration of the agent, improved binding of the CPNP to the membrane with resulting endocytosis, and the enhanced endosomal escape of the agent intracellularly. Whatever the mechanism may be, CPNPs are an especially non-toxic, easily synthesized delivery vehicle for gene vectors.
Initial attempts at gene therapy had the DNA coated on the outside of the nanoparticle [8, 9] as this was the simplest approach. The calcium is suspected to form a complex with the phosphate backbone of the DNA, and so the DNA quickly binds to the CPNP. However, this surface functionalization of the nanoparticle with DNA does not necessarily protect the exposed DNA from degradation by the in vivo environment. Therefore the DNA should be encapsulated by a shell of the calcium phosphate to provide further protection. One way in which this was done was by forming multi-shell particles in which the DNA was protected by a secondary shell of calcium phosphate[10]. Although effective in improving transfection, the multi-shell approach increases the size of the particles and their variance. Another method that we find promising is to co-precipitate the DNA and calcium phosphate via micro-emulsions.
While we believe the calcium phosphate nanoparticle will be an effective means of delivering the DNA origami into the cell, once the DNA is freely inside it still faces the challenges of avoiding intracellular enzyme degradation and passing the nucleus membrane barrier. There is little known about DNA origami within the cell. We do know that the structurally dense folded DNA can be resistant to DNase digestion of the structure for up to an hour[11] and structurally stable in cell lysate[12]. We also know of DNA origami being introduced intracellularly to provoke an immune response[13], but in general DNA origami has only been used as a vehicle for other agents. NO attempts have been made to design origami that delivers itself into the nucleus.
To design the origami to enter the cell requires that it be able to take advantage of the traffic of proteins entering the nucleus. In the non-dividing cell, the nucleus is only accessible by the nuclear pore complex, presenting a major barrier to importing the DNA. Nuclear targeting sequences can be used to enhance plasmid delivery, as these sequences can help form complexes with proteins that carry nuclear localization tags, which are responsible for chaperone delivery into the nucleus. The SV40 enhancer 72-base sequence is known to be especially effective in transporting plasmids into the nucleus[14], as it is suspscted to bind to transcription factors that are then imported. If the DNA origami were to be designed so that similar promoter/enhancer sequences were optimally located on the surface of the structure, then we would expect to see a significant increase in transfection in this regard as well.
Another potential advantage is the effect the DNA origami may have on the shape of the nanoparticle. DNA origami is most easily folded into cylindrical rods, and so the precipitation of the calcium phosphate onto the origami may induce a cylindrical shape to the nanoparticle as well. It was shown in 2008[15] that cylindrical nanoparticles with a high aspect ratio showed a higher rate of internalization than their more symmetrical counterparts. The potential mechanism may be reminiscent towards that of infectious rod-shaped prokaryotes, but that has yet to be determined.
See our Methods for more.
Our Focus
We first set out to develop a method of transfecting cells by a novel approach of encapsulating DNA origami with calcium phosphate to form biocompatible nanoparticles. This was a combinatorial process, in that both the DNA origami and calcium phosphate are well established but have never been combined in application. We hypothesized that the DNA origami will be well encapsulated by the calcium phosphate. If it is succesfully delivered into the cell, we hypothesize that the DNA origami will serve as an effective agent for gene transfection due to the enzyme resistant structure and potential for functional design.
We consider DNA origami to be the next step in transfection in tandem with calcium phosphate. This is because although calcium phosphate can show a high uptake efficiency, the transfection efficiency of the DNA is minimal ~1%. There needs to be a new delivery mechanism once the DNA is within the cell itself.
Feasibility
Calcium phosphate nanoparticle protocols will have to be optimized within the lab as the process is highly variable with the experimenter. Our lab is already established with DNA origami, and so the highest difficulty we will face is the imaging of the nanoparticles and origami, which can be technically arduous. We do not expect to complete phase 4 and demonstrate improved transfection within the nucleus because of time and technical constraints; we don't have the capability for the genetic engineering of the plasmid that would be necessary. Demonstrating an intact DNA origami structure within the cytosol is a good stepping off point because it validates our early arguments.
Economy
Most plasmid vectors in the market sell for several hundred dollars, and viral packages reach the several thousands. One limitation we face with the DNA origami technique is that the plasmid must be fully sequenced in order for the structure to be properly folded using the staple strand technique. However since most companies fully sequence their vector plasmids, the origami technique can be approximated for many commercial products. For future applications, we expect that as DNA synthesis and sequencing continues to become cheaper the cost of the DNA origami will be minimal in comparison to other techniques.
Our Goals
- Goal
- Goal
- Goal
- Gooooaal!
Resources
- Rothemund PW. Folding DNA to create nanoscale shapes and patterns. Nature. 2006 Mar 16;440(7082):297-302. DOI:10.1038/nature04586 |
- Douglas SM, Bachelet I, and Church GM. A logic-gated nanorobot for targeted transport of molecular payloads. Science. 2012 Feb 17;335(6070):831-4. DOI:10.1126/science.1214081 |
- Schüller VJ, Heidegger S, Sandholzer N, Nickels PC, Suhartha NA, Endres S, Bourquin C, and Liedl T. Cellular immunostimulation by CpG-sequence-coated DNA origami structures. ACS Nano. 2011 Dec 27;5(12):9696-702. DOI:10.1021/nn203161y |
- Lee H, Lytton-Jean AK, Chen Y, Love KT, Park AI, Karagiannis ED, Sehgal A, Querbes W, Zurenko CS, Jayaraman M, Peng CG, Charisse K, Borodovsky A, Manoharan M, Donahoe JS, Truelove J, Nahrendorf M, Langer R, and Anderson DG. Molecularly self-assembled nucleic acid nanoparticles for targeted in vivo siRNA delivery. Nat Nanotechnol. 2012 Jun 3;7(6):389-93. DOI:10.1038/nnano.2012.73 |
- Graham FL and van der Eb AJ. A new technique for the assay of infectivity of human adenovirus 5 DNA. Virology. 1973 Apr;52(2):456-67. DOI:10.1016/0042-6822(73)90341-3 |
- Altinoğlu EI, Russin TJ, Kaiser JM, Barth BM, Eklund PC, Kester M, and Adair JH. Near-infrared emitting fluorophore-doped calcium phosphate nanoparticles for in vivo imaging of human breast cancer. ACS Nano. 2008 Oct 28;2(10):2075-84. DOI:10.1021/nn800448r |
- Sokolova V, Knuschke T, Kovtun A, Buer J, Epple M, and Westendorf AM. The use of calcium phosphate nanoparticles encapsulating Toll-like receptor ligands and the antigen hemagglutinin to induce dendritic cell maturation and T cell activation. Biomaterials. 2010 Jul;31(21):5627-33. DOI:10.1016/j.biomaterials.2010.03.067 |
- Zhang G, Liu T, Chen YH, Chen Y, Xu M, Peng J, Yu S, Yuan J, and Zhang X. Tissue specific cytotoxicity of colon cancer cells mediated by nanoparticle-delivered suicide gene in vitro and in vivo. Clin Cancer Res. 2009 Jan 1;15(1):201-7. DOI:10.1158/1078-0432.CCR-08-1094 |
- Liu T, Tang A, Zhang G, Chen Y, Zhang J, Peng S, and Cai Z. Calcium phosphate nanoparticles as a novel nonviral vector for efficient transfection of DNA in cancer gene therapy. Cancer Biother Radiopharm. 2005 Apr;20(2):141-9. DOI:10.1089/cbr.2005.20.141 |
- Sokolova VV, Radtke I, Heumann R, and Epple M. Effective transfection of cells with multi-shell calcium phosphate-DNA nanoparticles. Biomaterials. 2006 Jun;27(16):3147-53. DOI:10.1016/j.biomaterials.2005.12.030 |
- Castro CE, Kilchherr F, Kim DN, Shiao EL, Wauer T, Wortmann P, Bathe M, and Dietz H. A primer to scaffolded DNA origami. Nat Methods. 2011 Mar;8(3):221-9. DOI:10.1038/nmeth.1570 |
- Mei Q, Wei X, Su F, Liu Y, Youngbull C, Johnson R, Lindsay S, Yan H, and Meldrum D. Stability of DNA origami nanoarrays in cell lysate. Nano Lett. 2011 Apr 13;11(4):1477-82. DOI:10.1021/nl1040836 |
- Schüller VJ, Heidegger S, Sandholzer N, Nickels PC, Suhartha NA, Endres S, Bourquin C, and Liedl T. Cellular immunostimulation by CpG-sequence-coated DNA origami structures. ACS Nano. 2011 Dec 27;5(12):9696-702. DOI:10.1021/nn203161y |
- Vacik J, Dean BS, Zimmer WE, and Dean DA. Cell-specific nuclear import of plasmid DNA. Gene Ther. 1999 Jun;6(6):1006-14. DOI:10.1038/sj.gt.3300924 |
- Gratton SE, Ropp PA, Pohlhaus PD, Luft JC, Madden VJ, Napier ME, and DeSimone JM. The effect of particle design on cellular internalization pathways. Proc Natl Acad Sci U S A. 2008 Aug 19;105(33):11613-8. DOI:10.1073/pnas.0801763105 |
- Paul M Gruenbacher 12:42, 14 April 2013 (EDT):