Registry of Standard Biological Models/Basic Component Models/Inverter
From OpenWetWare
Jump to navigationJump to search
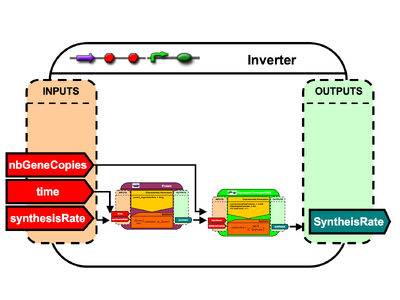
Inverter Architecture
CellML structure (CellML 1.1 spec)
- Component: Inverter
- Units:
- Already defined in sub-components
- Imports:
- Components:
- Repressed Promoter/RBS
- Protein
- Components:
- Variables:
- time (public interface = in)
- nbGeneCopies (public interface = in)
- protein (public interface = out / init value = 0.)
- MathML
- none
Comments
- No too sure how to connect the components together following the import
- Do I have to declare all the variables I need to use from the imported components ?
- How do I 'trigger' the dynamic behaviour of the sub-components ?