SBB10Ntbk-BenBubenheim
From OpenWetWare
Jump to navigationJump to search
My Project
My Personal Page
My Group Project Notebook
My Group Prompt
Construction Files
sbb01: N15 Protelomerase
Construction of sbb01: N15 Protelomerase
PCR BDBn15001F/BDBn15001R on Bca1455 (942 bp, gp=A)
PCR BDBn15002F/BDBn15002R on Bca1485 (1020 bp, gp=B)
--------------------------------------------------------------
PCR BDBn15001F/BDBn15002R on A+B (1924 bp, EcoR1/BamH1)
Sub in pBjk2741-Bca1144 (EcoRI/BamHI, 910+2170, L)
Product is pBjk2741-sbb01 {rbs.part!}
BDBn15001F Forward retrieval, RBS introduction and BioBricking on Bca1455
CCATAgaattcATGagatctTCAATCTCGGAGGAACGGTAAGGAAGGAAAACatgAGCAAGGTAAAAATCGGTGAG
BDBn15001R Reverse retrieval and SOE-site introduciton on Bca1455
CGCTTTTATCTTCACTGCGTTTTTTAGCTTGCCCTGAG
BDBn15002F Forward retrieval and SOE-site introduciton on Bca1485
CTCAGGGCAAGCTAAAAAACGCAGTGAAGATAAAAGCG
BDBn15002R Reverse retrieval and BioBricking on Bca1485
CTGATggatccGCTGTAGTACGTTTCCCATGCGG
JCA Notes
- got 974 for first pcr
- No stop codon on BDBn15002R
- Otherwise correct
- replaced first oligo with shorter, it's pressing your luck to use an oligo that long for soeing
- May need to alter this one based on PCR results
Construction of sbb01: N15 Protelomerase PCR BDBn15001F/CA1472 on Bca1455 1-3 (595bp, gp=A) PCR ca1461/BDBn15001R on Bca1455 1-1 (762bp, gp=B) PCR BDBn15002F/BDBn15002R on Bca1485 2-1 (1023 bp, gp=C) -------------------------------------------------------------- PCR BDBn15001F/BDBn15002R on A+B+C (1950 bp, EcoR1/BamH1) Sub in pBjk2741-Bca1144 (EcoRI/BamHI, 910+2170, L) Product is pBjk2741-sbb01 {rbs.part!} BDBn15001F Forward retrieval, RBS introduction and BioBricking on Bca1455 CCATAgaattcATGagatctGAGGAACGGTAAGGAAGGAAAACatgAGCAAGGTAAAAATCGG CA1461 GGAAAGGATTGCAGAAAAGAATAACCGCGAATACTTTTAACGCCTATATGA PCA assembly of protelomerasefiveprime (Bca1455) CA1472 TTGTTGGAATAACTTATAAAGATAGTCACGGCGCTCCTTCCAATCATCACT PCA assembly of protelomerasefiveprime (Bca1455) BDBn15001R Reverse retrieval and SOE-site introduciton on Bca1455 CGCTTTTATCTTCACTGCGTTTTTTAGCTTGCCCTGAG BDBn15002F Forward retrieval and SOE-site introduciton on Bca1485 CTCAGGGCAAGCTAAAAAACGCAGTGAAGATAAAAGCG BDBn15002R Reverse retrieval and BioBricking on Bca1485 CTGATggatccttaGCTGTAGTACGTTTCCCATGCGG part: gatctGAGGAACGGTAAGGAAGGAAAACatgAGCAAGGTAAAAATCGGTGAGTTGATCAACACGCTTGTGAATGAGGTAGAGGCCATTGATGCCTCAGACCGCCCACAAGGCGACAAAACGAAGAGAATTAAAGCCGCAGCCGCACGGTATAAGAACGCGTTATTTAATGATAAAAGAAAGTTCCGTGGGAAAGGATTGCAGAAAAGAATAACCGCGAATACTTTTAACGCCTATATGAGCAGGGCAAGAAAGCGGTTTGATGATAAATTACATCATAGCTTTGATAAAAATATTAATAAATTATCGGAAAAGTATCCTCTTTACAGCGAAGAATTATCTTCATGGCTTTCTATGCCTACGGCTAATATTCGCCAGCACATGTCATCGTTACAATCTAAATTGAAAGAAATAATGCCGCTTGCCGAAGAGTTATCAAATGTAAGAATAGGCTCTAAAGGCAGTGATGCAAAAATAGCAAGACTAATAAAAAAATATCCAGATTGGAGTTTTGCTCTTAGTGATTTAAACAGTGATGATTGGAAGGAGCGCCGTGACTATCTTTATAAGTTATTCCAACAAGGCTCTGCGTTGTTAGAAGAACTACACCAGCTCAAGGTCAACCATGAGGTTCTGTACCATCTCCAGCTAAGCCCTGCGGAGCGTACATCTATACAGCAACGATGGGCCGATGTTCTGCGCGAGAAGAAGCGTAATGTTGTGGTTATTGACTACCCAACATACATGCAGTCTATCTATGATATTTTGAATAATCCTGCGACTTTATTTAGTTTAAACACTCGTTCTGGAATGGCACCTTTGGCCTTTGCTCTGGCTGCGGTATCAGGGCGAAGAATGATTGAGATAATGTTTCAGGGTGAATTTGCCGTTTCAGGAAAGTATACGGTTAATTTCTCAGGGCAAGCTAAAAAACGCAGTGAAGATAAAAGCGTAACCAGAACGATTTATACTTTATGCGAAGCAAAATTATTCGTTGAATTATTAACAGAATTGCGTTCTTGCTCTGCTGCATCTGATTTCGATGAGGTTGTTAAAGGATATGGAAAGGATGATACAAGGTCTGAGAACGGCAGGATAAATGCTATTTTAGCAAAAGCATTTAACCCTTGGGTTAAATCATTTTTCGGCGATGACCGTCGTGTTTATAAAGATAGCCGCGCTATTTACGCTCGCATCGCTTATGAGATGTTCTTCCGCGTCGATCCACGGTGGAAAAACGTCGACGAGGATGTGTTCTTCATGGAGATTCTCGGACACGACGATGAGAACACCCAGCTGCACTATAAGCAGTTCAAGCTGGCCAACTTTTCCAGAACCTGGCGACCGGAAGTTGGGGATGAAAACACCAGGCTGGTGGCTCTCCAGAAACTGGACGATGAAATGCCAGGCTTTGCCAGAGGTGACGCTGGCGTCCGTCTCCATGAAACCGTTAAGCAGCTGGTGGAGCAGGACCCATCAGCAAAAATAACCAACAGCACTCTCCGGGCCTTTAAATTTAGCCCGACGATGATTAGCCGGTACCTGGAGTTTGCCGCTGATGCATTGGGGCAGTTCGTTGGCGAGAACGGGCAGTGGCAGCTCAAGATAGAGACACCTGCAATCGTCCTGCCTGATGAAGAATCCGTTGAGACCATCGACGAACCGGATGATGAGTCCCAAGACGACGAGCTGGATGAAGATGAAATTGAGCTCGACGAGGGTGGCGGCGATGAACCAACCGAAGAGGAAGGGCCAGAAGAACATCAGCCAACTGCTCTAAAACCCGTCTTCAAGCCTGCAAAAAATAACGGGGACGGAACGTACAAGATAGAGTTTGAATACGATGGAAAGCATTATGCCTGGTCCGGCCCCGCCGATAGCCCTATGGCCGCAATGCGATCCGCATGGGAAACGTACTACAGCTAAG
sbb05: piggieBac 5'TR
Construction of sbb05: piggieBac 5'TR Pool BDBpb001 through BDBpb010, assemble by PCA PCR BDBpb001/BDBpb010 on PCA reaction (344 bp, EcoRI/BamHI) Sub in pBjk2741-Bca1144 (EcoRI/BamHI, 910+2170, L) Product is pBjk2741-sbb05 ------------ BDBpb001 PCA assembly of sbb11 CCATAGAATTCATGAGATCTTTAACCCTAGAAAGATAGTCTGCGTAAAATTGACGC BDBpb002 PCA assembly of sbb11 GCTATTTAGAAAGAGAGAGCAATATTTCAAGAATGCATGCGTCAATTTTACGCAGACTATC BDBpb003 PCA assembly of sbb11 GAAATATTGCTCTCTCTTTCTAAATAGCGCGAATCCGTCGCTGTGCATTTAGG BDBpb004 PCA assembly of sbb11 CTCACGGGAGCTCCAAGCGGCGACTGAGATGTCCTAAATGCACAGCGACGGA BDBpb005 PCA assembly of sbb11 CGCTTGGAGCTCCCGTGAGGCGTGCTTGTCAATGCGGTAAGTGTCACTGATTTTGA BDBpb006 PCA assembly of sbb11 CGTCATTTTGACTCACGCGGTCGTTATAGTTCAAAATCAGTGACACTTACCGC BDBpb007 PCA assembly of sbb11 CCGCGTGAGTCAAAATGACGCATGATTATCTTTTACGTGACTTTTAAGATTTAACTCATACG BDBpb008 PCA assembly of sbb11 TATCACGTAAGTAGAACATGAAATAACAATATAATTATCGTATGAGTTAAATCTTAAAAGTCACGT BDBpb009 PCA assembly of sbb11 ATATTGTTATTTCATGTTCTACTTACGTGATAACTTATTATATATATATTTTCTTGTTATAGATATCGGATC BDBpb010 PCA assembly of sbb11 CTGATGGATCCGATATCTATAACAAGAAAATATATATATAATAA
JCA Notes
- Correct
Part Assembly
2/17/2010
sbb01
- Diluted original oligo stock from 100uM to 10uM (in separate tubes) as required by the protocol.
- Set up 3 PCR's as per Cloning by PCR with oligos:
- BDBn15001F/CA1472 on Bca1455 1-3 (A)
- ca1461/BDBn15001R on Bca1455 1-1 (B)
- BDBn15002F/BDBn15002R on Bca1485 2-1 (C)
24uL ddH2O 3.3uL 10x Expand Buffer "2" 3.3uL dNTPs (2mM in each) 1uL Oligo 1, 10uM 1uL Oligo 2, 10uM 0.5uL Expand polymerase "1" 0.5uL Template DNA
- Left labeled tubes for GSI's to put into the thermocycler for 2k55 program
sbb05
- Set up PCA assembly step as per PCA Gene Synthesis with the oligos in the sbb05 construction file
PCA assembly protocol 1. 38 uL ddH2O 2. 5 ul 10x expand buffer 3. 5 ul 2mM dNTPs 4. 1 ul oligo mixture (100uM total, mixture of oligos after combination of 100uM stocks) 5. 0.75 ul Expand polymerase Program (can run JCA/PCA1) 1. 2 min initial denature at 94oC 2. 30 sec denature at 94oC 3. 30 sec anneal at 55oC [This should be the overlap temp of your oligos - vary as needed] 4. 30 sec extension at 72oC 5. repeat 2-4 30 times total
- GSI's ran thermocycler
2/22/2010
sbb01
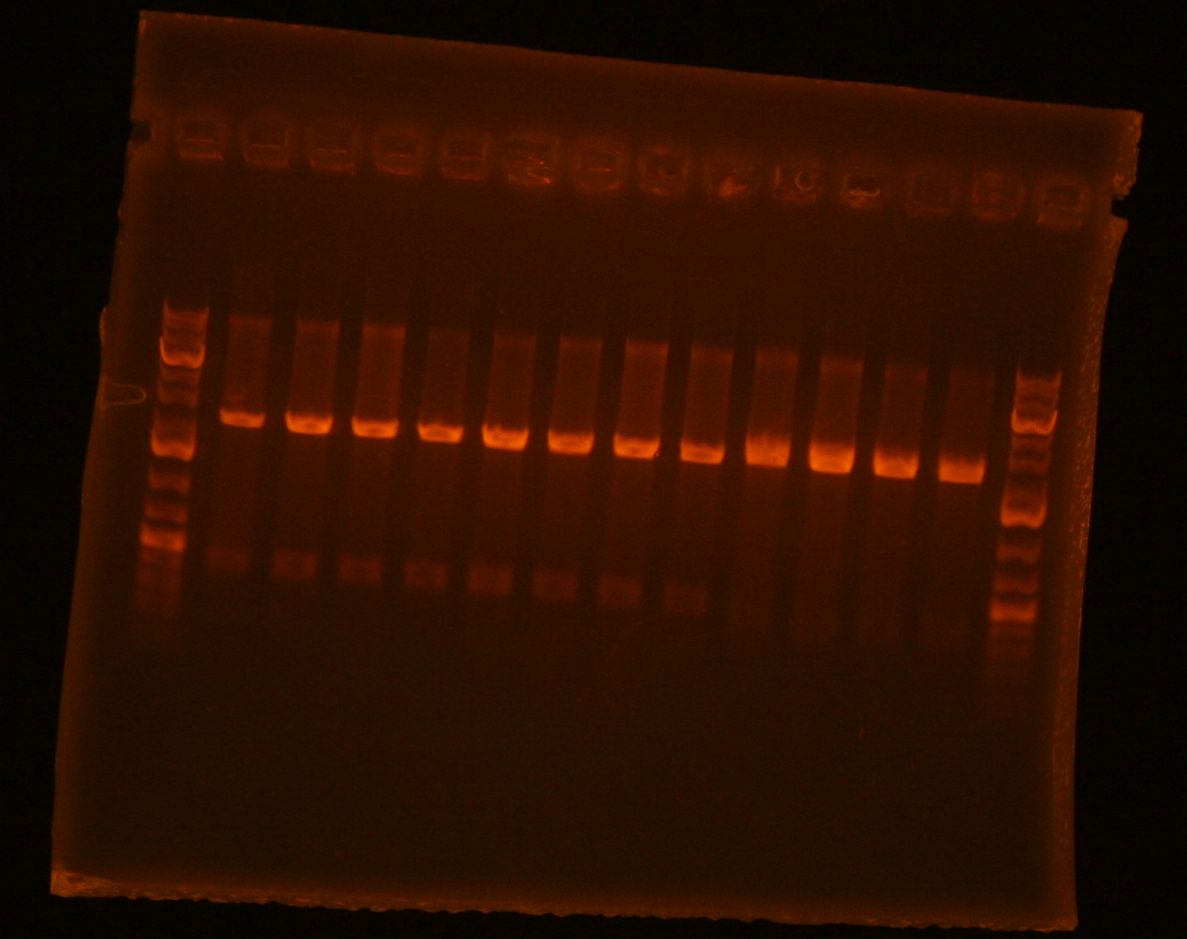
- set up PCR products A, B, and C for preparative gel.
- 6uL PCR product + 4uL loading buffer
- The gel with my samples was run by Amy.
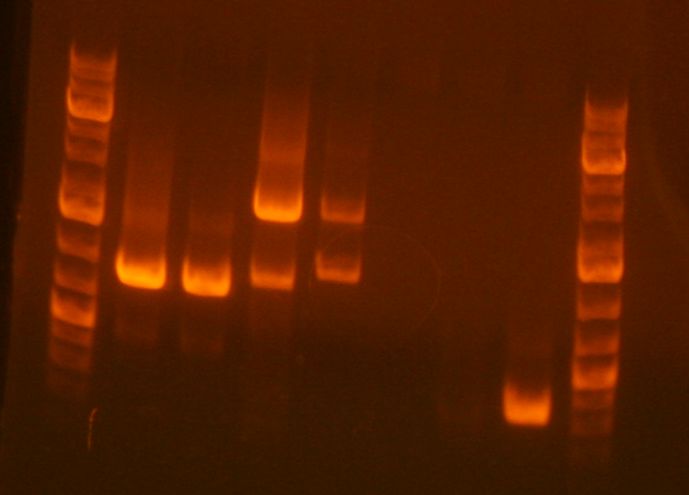
- My PCR products (lanes 7, 8 , 9) appear to be the correct sizes.
- GSI's cut out my bands and melted them in 650uL ADB buffer as per the first step in SOEing PCR
sbb05
- Performed Small-frag Zymo Cleanup to purify PCA Assembly.
1. Add 100 uL of Zymo ADB buffer (brown bottle) to the reaction. 2. Transfer into the Zymo column (small clear guys) 3. Add 500uL of Ethanol and pipette up and down to mix 4. spin through, discard waste. 5. Add 200 uL of PE or Zymo Wash buffer (which is basically 70% ethanol) 6. spin through, discard waste. 7. Add 200 uL of PE or Zymo Wash buffer 8. spin through, discard waste. 9. spin for 90 seconds, full speed to dry. 10. elute with water into a fresh Eppendorf tube
- With eluted product, set up amplification PCR
Recipe 1. 1 ul each outer oligo (10 uM) 2. .5 ul phusion 3. 10 ul 5x phusion buffer 4. 5 ul 2mM dNTPs 5. 32.5 ul H2O Program 1. 2 min initial denature at 94oC 2. 30 sec denature at 94oC 3. 30 sec anneal at 60oC [This should be high, as your outer oligos now have a huge overlap with the correct product] 4. 30 sec extension at 68oC 5. repeat 2-4 30 times total
- GSI's ran PCR program
2/24/2010
sbb01
- Purified PCR product mixture (A+B+C) by Zymo Gel Purification and eluted in 50uL ddH2O:
All spins until the drying step are 15 second full speed spins. 1. cut out bands minimizing extra gel matter. 2. put in ependorf tube and add 600uL of Zymo ADB buffer (brown bottle). 3. heat at 55, shake and/or vortex until the gel has dissolved. 4. If the DNA is <300bp add 250uL of isopropanol 5. transfer into the Zymo column inside a collection tube (small clear guys) 6. spin through, discard waste. 7. Add 200 uL of PE buffer (which is basically 70% ethanol) 8. spin through, discard waste. 9. Add 200 uL of PE buffer 10. spin through, discard waste. 11. spin for 90 seconds, full speed to dry. 12. elute with 8.5 uL of water into a fresh Eppendorf tube
- Set up second PCR step ( Cloning by PCR):
24uL ddH2O 3.3uL 10x Expand Buffer "2" 3.3uL dNTPs (2mM in each) 1uL Oligo 1, 10uM 1uL Oligo 2, 10uM 0.5uL Expand polymerase "1" 0.5uL Template DNA
- 2K55 program was used again.
sbb05
- Amplification product was retrieved and prepared for an analytical gel.
- 3uL PCR product + 7uL loading buffer
- Michel ran this gel.
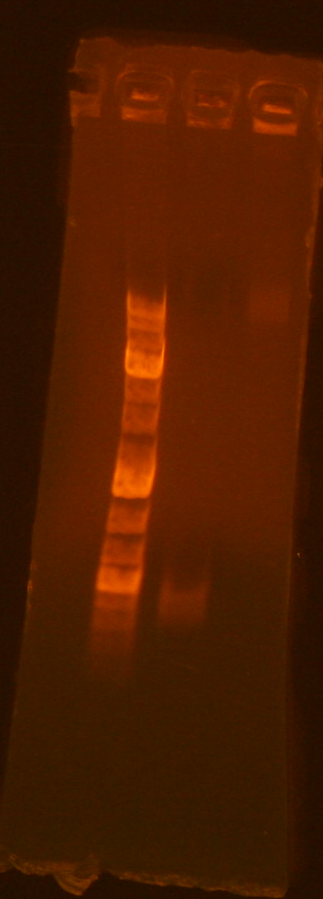
- My amplification product was in lane 2, and the band corresponded to the correct length.
3/1/2010
sbb01
- Prepared PCR product for analytical gel.
- 3uL PCR product + 7 uL loading buffer
- My sample is in the 5th lane. It looks good, although there is another band that looks like it is an incomplete fragment (A+B or B+C).
- Cleaned up the PCR product using a Regular Zymo Cleanup.
sbb05
- Cleaned up the PCA product using Regular Zymo Cleanup.
- Set up Eco/Bam digest for the eluted product:
8uL of eluted PCR product 1uL of NEB Buffer 2 0.5uL EcoRI 0.5uL BamHI
- Incubated at 37°C for an hour.
- Prepared digest for analytical gel
- 10 uL digest + 1 uL 6x Loading Buffer
- Ran the gel by myself, the band looks good
3/3/10
sbb01
- Prepared my PCR product for an Eco/Bam digestion by setting up the following reaction:
8uL of eluted PCR product 1uL of NEB Buffer 2 0.5uL EcoRI 0.5uL BamHI
- Incubated at 37°C for an hour.
- Prepared digest for preparative gel
- 10 uL digest + 1 uL 6x Loading Buffer
- My digest is in lane 3. It looks good.
- Cut out the gel and dropped it into an Eppendorf tube.
- Added 600 uL ADB buffer and melted the gel at 55°C
- Performed a Zymo Gel Purification:
* All spins until the drying step are 15 second full speed spins. 1. cut out bands minimizing extra gel matter. 2. put in ependorf tube and add 600uL of Zymo ADB buffer (brown bottle). 3. heat at 55, shake and/or vortex until the gel has dissolved. 4. If the DNA is <300bp add 250uL of isopropanol 5. transfer into the Zymo column inside a collection tube (small clear guys) 6. spin through, discard waste. 7. Add 200 uL of PE buffer (which is basically 70% ethanol) 8. spin through, discard waste. 9. Add 200 uL of PE buffer 10. spin through, discard waste. 11. spin for 90 seconds, full speed to dry. 12. elute with 8.5 uL of water into a fresh Eppendorf tube
sbb05
- Set up Ligation reaction as per Ligation Protocol with the pBjk2741 vector digest.
Set up the following reaction: 6.5uL ddH2O 1uL T4 DNA Ligase Buffer (small red or black-striped tubes) 1uL Vector digest 1uL Insert digest 0.5uL T4 DNA Ligase Pound upside down on the bench to mix Give it a quick spin to send it back to the bottom of the tube Incubate on the benchtop for 30min Put on ice and proceed to the transformation
- With ligated product, proceeded to transformation as per Transformation Protocol.
1. Thaw a 200 uL aliquot of cells on ice 2. Add 50 uL of water to the cells 3. Add 30 uL of KCM to the cells 4. Put your ligation mixture on ice, let cool a minute or two 5. Add 70 uL of the cell cocktail to the ligation, stir to mix 6. Let sit on ice for 10 min 7. Heat shock for 90 seconds at 42 (longer incubation may work better) 8. Put back on ice for 1 min 9. Add 100uL of 2YT, let shake in the 37 degree incubator for 1 hour 10. Plate 70+ uL on selective antibiotics, let incubate at 37 degrees overnight
3/8/10
sbb01
- Prepared a ligation reaction with the Eco/Bam digest of my sbb01 part and pBjk2741 as the vector.
- proceeded to transformation after the ligation (post-30min RT incubation) was left in ice for awhile while i finished the miniprep for sbb05
- GSI's plated my transformants as the recovery
sbb05
- GSI's picked 4 colonies labeled 1-4
- Proceeded with mini-prep as per Miniprep Protocol.
MINIPREP (1mL - 5mL) Procedure for Plasmid DNA Purification (using the QIAGEN QIAPrep Spin Miniprep kit) 1. Pellet around 1.5 mL or 2 mL saturated culture by spinning full speed, 30 seconds. 2. Dump supernatant, repeat to pellet another 1.5 mL (for a total of 3 mL) 3. Add 250uL of P1 buffer into each tube. Resuspend the cells using a vortexer. 4. Add 250uL of P2 buffer (a base that denatures everything and causes cells to lyse). Gently mix up and down. Solution should become clearer. 5. Add 350uL of N3 buffer (an acid of pH ~5 that causes cell junk - including protein and chromosomal DNA - to precipitate, and leaves plasmids and other small molecules in solution). Slowly invert a few times, then shake. 6. Spin in centrifuge at top speed for 5 minutes. 7. Label blue columns with an alcohol-resistant lab pen. 8. Pour liquid into columns, and place the columns into the centrifuge. Spin at 12000 rpm for 30 seconds. 9. Dump liquid out of the collectors under the columns (the DNA should be stuck to the white resin) 10. Wash each column with 500 uL of PB buffer. 11. Spin in centrifuge at 12000rpm for approximately 15 seconds, then flick out the liquid again. 12. Wash with 750uL of PE buffer (washes the salts off the resins). 13. Spin in centrifuge at 12000rpm for approximately 15 seconds and flick out liquid again. 14. Spin in centrifuge at full speed for 1 minute to dry off all water and ethanol. 15. Label new tubes and put columns in them. 16. Elute them by squirting 50uL of water down the middle of the column (don't let it stick to the sides). 17. Spin in centrifuge at top speed for 30 seconds. 18. Take out columns and cap the tubes. 19. Clean up - note the P1 buffer is stored at 4degC and all the rest at room temperature.
- Stored eluted product in tubes labeled 1-4
3/10/10
sbb01
- NOTE: the LB used to rescue last time turned out to be contaminated, plates have small white spots that may be fungi
- Mini-prepped the colonies that the GSI's picked as per Miniprep Protocol.
- saved in tubes for digestion and mapping next time
sbb05
- digested mini-prep elutions with eco and bam as per Mapping Digest.
4uL ddH2O 4uL Miniprepped plasmid 1uL 10x NEB Buffer 2 0.5uL EcoRI 0.5uL BamHI (for parts >250bp) or XhoI (for parts <250bp) Incubate at 37°C for 30min
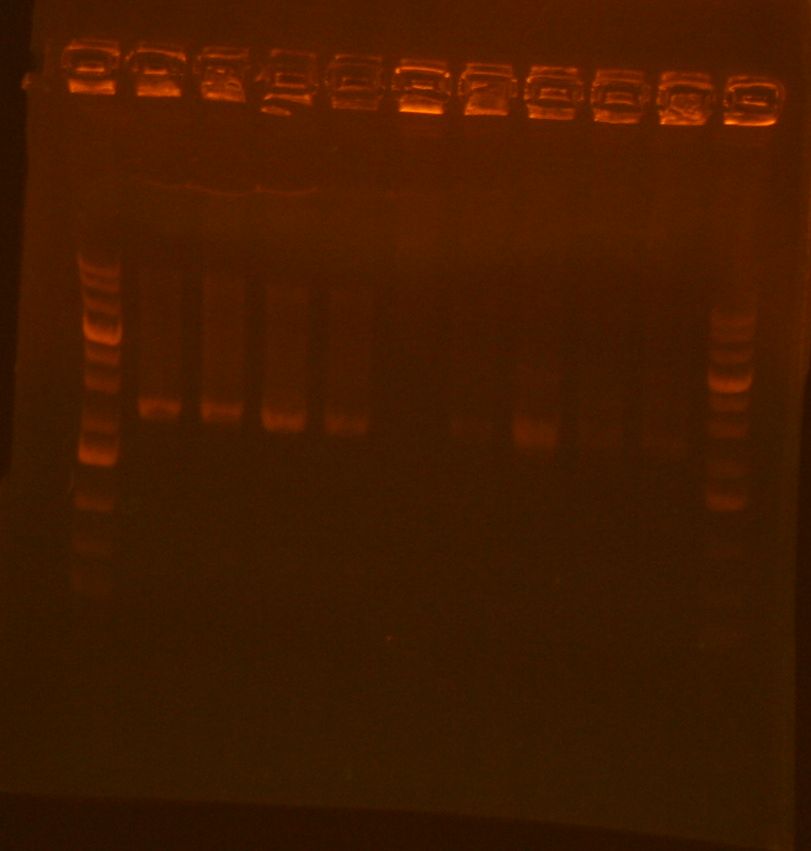
- added 1uL of 6x loading dye and ran analytical gel
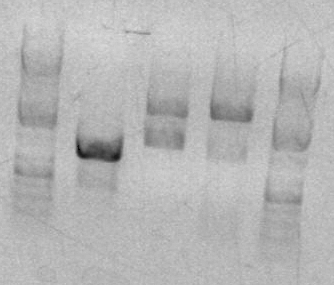
- had to re-stain the gel, but bands showed up around 344bp which is good.
- my lanes are the rightmost 4, had to take a picture without the plate underneath to see the bands
- submitted all 4 clones for sequencing
3/15/10
sbb01
- digested with EcoRI/BamHI according to Mapping Digest.
- my lanes are the rightmost 4, band around 2000 doesn't exactly show that my part is there, but since there is a line at 2000 and not 4000 it shows that the plasmid was double digested so my part should be in there, will send to sequencing.
sbb05
- recieved sequencing results:
rev-comp s-w opt: 1565 Z-score: 2302.4 bits: 436.2 E(): 2e-126
Smith-Waterman score: 1565; 100.000% identity (100.000% ungapped) in 313 nt overlap (883-1195:1-313)
860 870 880 890 900 910
Seq1- TAAGGATCTGAAGTGGAATTCATGAGATCTTTAACCCTAGAAAGATAGTCTGCGTAAAAT
::::::::::::::::::::::::::::::
Seq2 TTAACCCTAGAAAGATAGTCTGCGTAAAAT
10 20 30
920 930 940 950 960 970
Seq1- TGACGCATGCATTCTTGAAATATTGCTCTCTCTTTCTAAATAGCGCGAATCCGTCGCTGT
::::::::::::::::::::::::::::::::::::::::::::::::::::::::::::
Seq2 TGACGCATGCATTCTTGAAATATTGCTCTCTCTTTCTAAATAGCGCGAATCCGTCGCTGT
40 50 60 70 80 90
980 990 1000 1010 1020 1030
Seq1- GCATTTAGGACATCTCAGTCGCCGCTTGGAGCTCCCGTGAGGCGTGCTTGTCAATGCGGT
::::::::::::::::::::::::::::::::::::::::::::::::::::::::::::
Seq2 GCATTTAGGACATCTCAGTCGCCGCTTGGAGCTCCCGTGAGGCGTGCTTGTCAATGCGGT
100 110 120 130 140 150
1040 1050 1060 1070 1080 1090
Seq1- AAGTGTCACTGATTTTGAACTATAACGACCGCGTGAGTCAAAATGACGCATGATTATCTT
::::::::::::::::::::::::::::::::::::::::::::::::::::::::::::
Seq2 AAGTGTCACTGATTTTGAACTATAACGACCGCGTGAGTCAAAATGACGCATGATTATCTT
160 170 180 190 200 210
1100 1110 1120 1130 1140 1150
Seq1- TTACGTGACTTTTAAGATTTAACTCATACGATAATTATATTGTTATTTCATGTTCTACTT
::::::::::::::::::::::::::::::::::::::::::::::::::::::::::::
Seq2 TTACGTGACTTTTAAGATTTAACTCATACGATAATTATATTGTTATTTCATGTTCTACTT
220 230 240 250 260 270
1160 1170 1180 1190 1200 1210
Seq1- ACGTGATAACTTATTATATATATATTTTCTTGTTATAGATATCGGATCCTAACTCGCTCC
:::::::::::::::::::::::::::::::::::::::::::
Seq2 ACGTGATAACTTATTATATATATATTTTCTTGTTATAGATATC
280 290 300 310
1220 1230 1240 1250 1260
Seq1- TCAGGCTTCCTCGCTCACTGACTCGCTGCGCTCGGTCTTCGGCGGGGCTGCC
- 100% alignment, sequence was seq1 and target was seq2, it worked!