User:Perry/Summer 2006 Harvard iGEM work
6/12/06
Incubated DNA nanostructure folding interactions. DNA scaffold and oligonucleotides designed by Shawn Douglas. Protocol available here. Included 2 negative controls, one with water instead of scaffold, one with water instead of oligonucleotides.
Transformed R0010, E0241, and E7104 parts into 15 ul aliquots of chemically competent OneShot Top10 cells.
6/13/06
Ran gels from nanostructure incubations yesterday. 2% agarose gel, 130V, 45min. Loaded 10ul sample + 1ul dye, and 10ul 1kb ladder.
Lane 5: 1 kb ladder
Lane 6: DNA nanostructure (+)
Lane 7: (-) control with no oligonucleotides
Lane 8: (-) control with no DNA scaffold
Prepared 2 liquid cultures from each transformation, 5ml LB + 50ul of 5mg/ml ampicillin, shaken at 180rpm and 37dC overnight.
6/14/06
Retrieved liquid cultures (3x2). I accidentally dropped culture #2 from R0010 transf and spilled half the volume. Set aside 1ml of each culture for glycerol stock. Performed miniprep on remaining culture. Accidentally performed 2 washes with PE. Eluted with 50ul water.
The nanodrop gave negative readings of DNA concentration for the minipreps.
Prepared 25ul digest reactions with 8ul of both minipreps of R0010 (SpeI/PstI) and of E0241 (XbaI/PstI). Incubated 1 hour at 37dC held at 4dC, then incubated at 80dC for 20 min. Added 2ul loading dye to each digest mixture, then ran total mixtures in gel, 45 min, 130V.
Lane 8: 1 kb ladder
Lane 9: R0010, miniprep from culture #1, SpeI/PstI digested
Lane 10: R0010, miniprep from culture #2, SpeI/PstI digested
Lane 11: E0241, miniprep from culture #1, XbaI/PstI digested
Lane 12: E0241, miniprep from culture #2, XbaI/PstI digested
We got no bands, and so performed no gel extraction. We probably had a failed miniprep, evident from the negative number Lewis got from the nanodrop. I guess 2 PE washes is bad.
6/15/06
Using Tiffany and Matt's gel extracted vector/insert, we took 2ul of their vector and 6ul of their insert and performed a ligation. Then we transformed into Top10.
6/16/06
Observed two colonies on transformation plate which fluoresced green under microscope (probably due to leaky expression of GFP). Left plates in refrigeration.
6/19/06
Picked the two colonies from the transformation plate and prepared two 5ml inoculations with 50ul amp (5mg/ml).
6/20/06
Miniprepped two inoculations, eluting with 30ul water. Nanodrop read 98ng/ul for #1 and 73ng/ul for #2. We prepared 25ul XbaI/PstI digests of each, using 8ul DNA miniprep, incubating 90min 37dC, 15min 80dC. Then we ran the digests on a 1.2% e-gel, using 10ul digest + 10ul water. We also loaded 10ul 1X 1kb ladder + 10ul water.
Lane 1: 1kb+ ladder
Lane 5: XbaI/PstI digest of R0010-E0241 #1
Lane 6: XbaI/PstI digest of R0010-E0241 #2
6/22/06
Prepared sequencing reactions. Chris had ordered new primers, reconstituted at 20uM. I made a 1:10 dilution for 2uM.
PT001 8ul miniprep #1, 4ul VF2
PT002 8ul miniprep #1, 4ul VR
PT003 8ul miniprep #2, 4ul VF2
PT004 8ul miniprep #2, 4ul VR
7/3/06
Went to MIT to pick up plasmids with streptavidin clones from Mark Howarth at the Ting lab (Howarth et al, 2006).
- Streptavidin, wild-type, 0.1 ug/ul (StrepWT)
- Streptavidin, wild-type plus His6 tag (alive), 0.09 ug/ul (StrepH)
- Streptavidin, mutant (dead), 0.09 ug/ul (StrepD)
7/5/06
Ordered primers to PCR out the streptavidin clones.
Diluted plasmids 1:50, 1ul + 49ul water.
Transformed with OneShot Top10 chemically competent cells. Aliquotted 2x15ul for pUC19 (+) and for (-). Aliquotted 3x30ul for StrepWT, StrepH, and StrepD. Used 5ul of plasmid 1:50 dilutions and 2.5ul of pUC19. Put plates in incubator at 5:20pm.
7/6/06
Took plates out at 10:30am; they were overgrown. I had difficulty picking out single colonies. I streaked and inoculated liquid cultures from two colonies from each transformation. Liquid culture: 30ml LB + 300ul amp, 6x4.5ml aliquots. Put into incubation at 11:00am.
Returned at 1:00am to take out streak plate and to perform miniprep. Included PB step and eluted with 30ul water. The tubes for elution of minipreps from colonies StrepWT #1 and StrepH #2 cracked in the centrifuge. Prepared new 4.5ml liquid cultures from the streaked colonies; put in incubator around 2:00am.
7/7/06
Received StrepMF and StrepMR primers; reconstituted them in EB at 100uM. Then made another dilution to 5uM with 5ul of 100uM + 95ul EB. Did the same with OmpAF and OmpAR primers.
Nanodropped the StrepD minipreps, got a rather jagged curve. StrepD#1 was ~45ng/ul, StrepD#2 was ~25ng/ul. I made dilutions of 1:50 and 1:25 respectively in water.
Made a 1:2 dilution in water of the 1:50 dilution of original SA-dead (aka StrepD) from MIT (made 7/5/06), thus a final dilution of 1:100.
Prepared three PCR reactions. I tried to go for 50ul total with 45ul Platinum PCR Supermix, 200nM each primer, 1ng template DNA.
(0) 45ul Platinum PCR Supermix + 2ul StrepMF (5uM) + 2ul StrepMR (5uM) + 1ul SA-dead (1:100)
(1) " + 1ul StrepD#1 (1:50)
(2) " + 1ul StrepD#2 (1:25)
Incubated 95dC for 10min, 30 x (95dC for 1min, 55dC for 1min, 72dC for 1min), 72dC for 10min, 4dC forever.
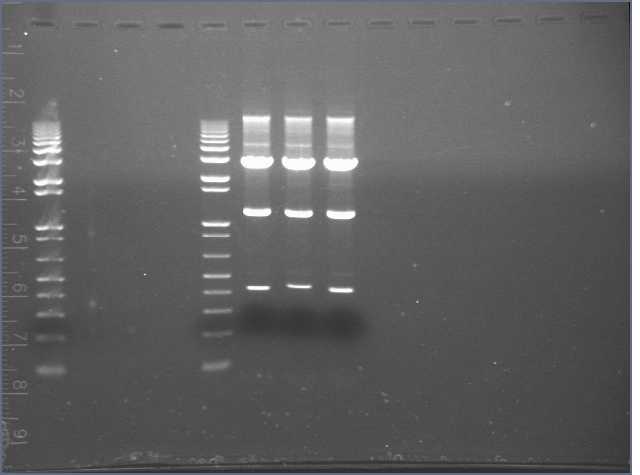
Ran PCR reactions on 1.2% e-gel for 15min. 10ul of each + 10ul water, 10ul 1X 1kb ladder + 10ul water. Included a lane of 10ul 5uM StrepMR primer + 10ul water to test visualization of ssDNA with ethidium bromide.
Lane 1: 1kb ladder
Lane 2: PCR (0)
Lane 3: PCR (1)
Lane 4: PCR (2)
Lane 5: 1kb ladder
Lane 6: StrepMR primer
The StrepMR primer showed a visible band.
None of the PCR reactions were successful. After brief contemplation, I realized a very good reason why. StrepMF and StrepMR were oligos designed for mutagenesis and NOT as forward/reverse primers for PCRing the mutant StrepD.
However, it's confusing that there isn't even a primer dimer band. StrepMF/StrepMR are complementary over ~15 base pairs.
Removed liquid cultures (StrepWT#1, StrepH#2) from last night at 1:30pm. Pelleted and left in freezer over lunch. Completed miniprep upon return. Included PB step and eluted with 30ul water.
7/10/06
Made a 1:2 dilution in water of the 1:50 dilution of original StrepWT and StrepH from MIT (made 7/5/06), thus a final dilution of 1:100.
Nanodropped the minipreps for ng/ul, and made corresponding dilutions in water. StrepWT#1, 41.2, 1:40. StrepWT#2, 23.9, 1:20. StrepH#1, 21.3, 1:20. StrepH#2, 53.7, 1:50. StrepD#1, 44.9, 1:40, StrepD#2, 15, 1:15.
Received primers StrepF, StrepR, StrepFS, StrepRS, StrepHR, StrepHRS. Reconstituted in EB to 100uM, and then diluted 10 of 100uM with 90ul EB for a 10uM diluted stock. Also made 10uM diluted stock from 100uM stocks StrepMF and StrepMR received 7/7/06.
Prepared PCR reactions. Made a partial master mix (enough for 10 tubes) of 450ul Platinum PCR Supermix + 10ul StrepF(10uM) + 10ul StrepMF(10uM) + 10ul StrepMR(10uM). Aliquotted 45ul to 9 tubes as labeled below. Then added 1ul reverse primer and 1ul template DNA as appropriate. I then realized that I should have aliquotted 48ul instead of 45ul, and added 3 more ul of partial master mix to each tube; however, I ended up with an empty tube when I should have had ~45ul extra. Something possibly went wrong in aliquotting?
All primers at 10uM. All template DNA ~1ng/ul.
W0: (45ul Platinum PCR Supermix + 1ul StrepF + 1ul StrepMF + 1ul StrepMR) + 1ul StrepRS + 1ul StrepWT plasmid (1:100 from MIT)
W1: ( ) + 1ul StrepRS + 1ul StrepWT#1 miniprep(1:40)
W2: ( ) + 1ul StrepRS + 1ul StrepWT#2 miniprep(1:20)
H0: ( ) + 1ul StrepHRS + 1ul StrepH plasmid (1:100 from MIT)
H1: ( ) + 1ul StrepHRS + 1ul StrepH#1 miniprep(1:20)
H2: ( ) + 1ul StrepHRS + 1ul StrepH#2 miniprep(1:50)
D0: ( ) + 1ul StrepRS + 1ul StrepD plasmid (1:100 from MIT)
D1: ( ) + 1ul StrepRS + 1ul StrepD#1 miniprep(1:40)
D2: ( ) + 1ul StrepRS + 1ul StrepD#2 miniprep(1:15)
Put into incubator with PCR program. Modified from an initial 95dC for 10min with hot start, to initial 95dC for 15min without hot start. (I got annoyed with having to go back to the PCR machine to make it continue from pause.)
Took 10ul from each PCR and of 1X 1kb+ ladder, mixed with 10ul water, and loaded into 1.2% e-gel.
Lane 1,11: 1kb+ ladder
Lanes 2-4: W0,1,2
Lanes 5-7: H0,1,2
Lanes 8-10: D0,1,2
Lane 12: Water
Ug, no bands of the expected 1kb size. There is a uniform band around 100bp, most probably a primer dimer.
Sigh, time for troubleshooting.
Prepared new 3ml liquid cultures from streak plates of W,H,D-1,2.
7/11/06
Received E. coli K12 genomic DNA (5ug) from ATCC. Rehydrated in 100ul water (for 50ng/ul) and incubated at 37dC for one hour, as directed.
Repeated nine PCRs from yesterday, but using 1ul of 1:50 plasmid from MIT and undiluted minipreps. Included one PCR reaction of 1:50 StrepW (plasmid from MIT) with water replacing the StrepMF and StrepMR primers, and one PCR reaction with 2ul OmpAF(5uM), 2ul OmpAR(5uM), and 2ul K12 genome(50ng/ul).
Ran e-gel, loading each well with 10ul PCR mixture/ladder + 10ul water.
Lane 1: 1kb+ ladder
Lanes 2-4: W0,1,2
Lanes 5-7: H0,1,2
Lanes 8-10: D0,1,2
Lane 11: W0 without StrepMF/StrepMR
Lane 12: OmpA
The Strep bands are more significant. Same primer dimers around 100, but then there's an expected weak band in lanes 2-10 at around 400, but a strong band in lane 11. There's also another weak band around 300. I believe that the mutagenesis primers are screwing up the efficiency of the PCR. There's no visible band for OmpA in lane 12.
Performed PCR purification of W0 without StrepMF/StrepMR and of OmpA PCRs.
Performed agarose gel electrophoresis of W,H,D-0,1,2 (poor loading of lanes 3 and 7).
Lanes 1,11: 1kb+ ladder
Lanes 2-4: W0,1,2
Lanes 5-7: H0,1,2
Lanes 8-10: D0,1,2
I forgot to take a picture before I excised 400bp bands from W2, H2, and D1 and disposed of the gel.
Prepared PCR reactions of W,H,D0 without StrepMF/R. Also prepared a 2nd PCR reaction with 1ul PCR-purified OmpA, and a 1st PCR reaction for OmpA using 10ul of K12 genomic DNA (50ng/ul, so 500ng), filling to a 60ul PCR mixture, with 2.4ul of OmpAF and OmpAR (5uM).
Removed liquid cultures of W,H,D-1,2 and refrigerated.
7/12/06
Performed gel purification of W2, H2, D1 gel slices from yesterday. Have they been successfully mutated to lose the XbaI site? We'll have to see.
Ran 1.2% e-gel of PCRs.
Lane 1: 1kb+
Lane 2,3,4: W0,H0,D0 without StrepMF/R
Lane 5: OmpA (a 2nd PCR of PCR-purified OmpA)
Lane 6: OmpA (a 1st PCR of 500ng K12 genomic DNA)
Performed PCR purifications of W0,H0,D0 without StrepMF/R and OmpA (2nd PCR).
Performed TOPO cloning with 4ul of gel-purified W2,H2,D1 (undetermined mutagenesis) and PCR-purified W0,H0,D0 (no mutagenesis) and OmpA (2nd PCR), 1ul salt solution, 1ul TOPO vector. Transformed each with 20ul Top10 chemically competent cells, including a (+) with 2.5ul pUC19 and a (-) with nothing.
I talked to Chris and realized that I've been going about mutagenesis completely wrong-ly. So I redid the mutageneses right-ly. Prepared six PCR reactions (all primers 10uM):
- 47ul Platinum PCR Supermix + 1ul StrepF + 1ul StrepMR + 1ul StrepW,H,D (1:50 plasmids from MIT)
- " + 1ul StrepRS + 1ul StrepMF + 1ul StrepW,D
- " + 1ul StrepHRS + 1ul StrepMF + 1ul StrepH
PCR purified the mixtures. Then using those, I prepared three PCRs
- StrepW(mut) = 45ul Platinum PCR Supermix + 2ul StrepW(F/MR) + 2ul StrepW(RS,MF)
- StrepH(mut) = " + 2ul StrepH(F/MR) + 2ul StrepH(HRS,MF)
- StrepD(mut) = " + 2ul StrepD(F/MR) + 2ul StrepD(RS,MF)
After 10/30 cycles, I paused the PCR machine and added 1ul StrepF and 1ul StrepRS to StrepW and StrepD, 1ul StrepF and 1ul StrepHRS to StrepH. Then I restarted the PCR machine to finish the remaining 20 cycles.
7/13/06
I took out the plates from the TOPO cloning/transformation; got a lot of colonies on all plates except (-) which had no colonies. I threw out the plates transformed with TOPO clone of gel-purified W1,H1,D2, and I threw out those gel-purified samples as well (because they had come from an incorrect PCR in which I mixed both forward/reverse and mutagenesis primers).
I streaked and inoculated 3ml cultures (30ul of 5mg/ml ampicillin) from the StrepW,H,D (no mut) and full OmpA transformations. Put in incubation around 2:00pm.
PCR purified StrepW,H,D(mut) from yesterday.
Ran 1.2% e-gel of the PCR purifications from yesterday and today.
Lanes 1,11: 1kb+ ladder
Lanes 2-4: StrepW(F/MR),(RS/MF),(mut)
Lanes 5-7: StrepH(F/MR),(HRS/MF),(mut)
Lanes 8-10: StrepD(F/MR),(RS/MF),(mut)
Topo cloned/transformed StrepW,H,D(mut). Aliquotted 15ul of Top10 chemically competent cells for the transformations, 7ul for (+) and 7ul for (-). Put in incubation around 2:00pm.
7/14/06
Took StrepW,H,D(no mut) and full OmpA liquid cultures out at 8:00am. Pelleted.
Took StrepW,H,D(mut) plates out; got a lot of colonies. Streaked and inoculated 3ml liquid cultures around 8:30am.
Miniprepped StrepW,H,D(no mut) and (mut), and full OmpA. Included PB step and eluted with 40ul water.
Prepared seven 20ul digests. First made a master mix of 20ul Buffer2, 20ul water, 4ul XbaI, 4ul PstI, 2ul BSA(100X), enough for 10. Aliquotted 5ul of master mix to each tube, then added 15ul of miniprep to each tube. Incubated at 37dC for 12h, 80dC for 20min, 4dC forever.
Received single-chain dimeric streptavidin from Dr. Filiz Aslan, as dry samples in tubes: SCD-NM (not mutant), C2, E2. She told me there was about 100ng in each tube. Reconstituted in 50ul EB for 2ng/ul, then made a 20ul stock, 1:2, for 1ng/ul.
Transformed SCD-NM, C2, E2, using 20ul Top10 OneShot chemically competent cells and 1ul of DNA (1ng/ul). Left in incubation at 1:30am.
7/15/06
Took out transformation plates; got a lot of colonies. Streaked and inoculated 3ml cultures from one colony from each plate: SCD-NM, C2, E2, at around 4:30pm.
Ran XbaI/PstI digests on 1% agarose gel.
Lane 1: 1kb+ ladder
Lanes 3,7,11: StrepW,H,D(no mut)
Lanes 5,9,13: StrepW,H,D(mut)
Lane 15: full OmpA
The 300bp/120bp bands in lanes 3, 7, 11 are the segments of the streptavidin BioBrick which have been cleaved by XbaI because of the XbaI site at bp#120 in the sequence. The 400bp band in lanes 5, 9, 13 is the full streptavidin BioBrick, not cleaved by XbaI because of the mutagenesis removing the XbaI restriction site. The 50bp band in all the lanes is the segment of the TOPO vector from the BioBrick PstI site to an XbaI site 50bp downstream. The 3kb/1kb bands in all the lanes are the segments of the rest of the TOPO vector, cleaved by XbaI, I think.
Excised the 400bp bands from lanes 5, 9, 13, and 1kb band from lane 15.
I just realized the futility of excising the OmpA 1kb band. First, that band is mixed with the 1kb band from the Topo vector (there is an XbaI site 1kb upstream of the insertion site). Second, I don't need a digested OmpA because I will not be using the full version.
7/16/06
Miniprepped SCD-NM, C2, E2 at around 8:30am. Included PB step and eluted with 30ul water.
7/17/06
Rec'd primers and reconstituted all in EB to 100uM, then made a dilution of 5ul of 100uM + 95ul EB, for 5uM.
Prepared 11 PCR reactions. Used SCD-NM, C2, E2 2ng/ul reconstitutions from 7/15/06, OmpA miniprep from 7/14/06, and 5uM primers. Also prepared 1 PCR reaction to check for OmpA insertion into the Topo vector using M13 forward and reverse primers at 20uM.
S, C, E(F/R): 45ul Platinum Supermix HiFi + 2ul StrepSCDF + 2ul StrepSCDRS + 1ul S, C, E
S, C, E(F/MR): " + 2ul StrepSCDF + 2ul StrepSCDMR + 1ul S, C, E
S, C, E(MF/R): " + 2ul StrepSCDMF + 2ul StrepSCDRS + 1ul S, C, E
O66: " + 2ul OmpA46F + 2ul OmpA66R + 1ul OmpA
O159: " + 2ul OmpA46F + 2ul OmpA159R + 1ul OmpA
OM13: " + 0.5ul M13F + 0.5ul M13R + 3ul water + 1ul OmpA
Sent out sequencing reactions to Genewiz, using the StrepW,H,D(mut) and OmpA minipreps from 7/14/06 (depleted the supply).
PT005: 8ul StrepW(mut), M13F(-21) to be added
PT006: 8ul StrepW(mut), M13R to be added
PT007: 8ul StrepH(mut), M13F(-21) to be added
PT008: 8ul StrepH(mut), M13R to be added
PT009: 8ul StrepD(mut), M13F(-21) to be added
PT010: 8ul StrepD(mut), M13R to be added
PT011: 8ul OmpA, M13F(-21) to be added
PT012: 8ul OmpA, M13R to be added
Ran a 1% agarose gel of the PCR reactions. Added 5ul loading dye, then loaded 40ul into wells.
Lane 1, 14, 15: 1kb+ (or at least I thought)
Lanes 2-4: SCD-NM MF/R, F/MR, F/R
Lanes 5-7: C2 MF/R, F/MR, F/R
Lanes 8-10: E2 MF/R, F/MR, F/R
Lane 11: OmpA ompA46F/ompA66R
Lane 12: OmpA ompA46F/ompA159R
Lane 13: OmpA M13F(-20)/M13R
Excised ~300bp band from lanes 2, 5, 8; ~800bp band from lanes 3, 6, 9; ~1kb band from lanes 4, 7, 10; ~150bp band from lane 11; ~400bp band from lane 12.
Performed gel extraction. Included extra QG wash and eluted with 30ul EB.
Ran a PCR with 45ul PCR Supermix HiFi + 2ul S(MF/R) gel extraction + 2ul S(F/MR), " + C(MF/R) + C(F/MR), " + E(MF/R) + E(F/MR). After cycle 10, I added 2ul StrepSCDF (5uM) and 2ul StrepSCDRS (5uM) to each reaction.
7/18/06
Rec'd LppFS, Lpp29R, Lpp78R, Lpp78RS primers. Reconstituted in EB to 100uM, then made a dilution to 5uM.
Prepared three PCR reactions, primers at 5uM.
Lpp29: 45ul Platinum PCR Supermix HiFi + 2ul K12 genomic DNA (50ng/ul) + 2ul LppFS + 2ul Lpp29R
Lpp78: " + " + " + 2ul Lpp78R
Lpp78S: " + " + " + 2ul Lpp78RS
Ran 1% agarose gel of PCRs (made gel larger than usual, 120ml TBE and 1.2g agarose). Added 5ul loading dye to each reaction and loaded 40ul. 130V, 50min.
Lane 1: 1kb+
Lane 2: SCD-NM(mut)
Lane 3: C2(mut)
Lane 4: E2(mut)
Lane 5: Lpp29
Lane 6: Lpp78
Lane 7: Lpp78S
Cut out bands from lanes 5, 6, 7. Gel extracted Lpp29 and Lpp78S.
Prepared six PCRs using yesterday's gel-extracted S, C, E (F/R) as template.
45ul Platinum PCR Supermix HiFi + 2ul StrepSCDMF + 2ul StrepSCDRS + 1ul S,C,E(F/R)
45ul Platinum PCR Supermix HiFi + 2ul StrepSCDF + 2ul StrepSCDMR + 1ul S,C,E(F/R)
Ran PCR reactions on 120ml 1% agarose gel (added 5ul loading dye, loaded 40ul). 130V, 50min
Lane 1: 1kb+
Lanes 2,4,6: S, C, E (MF/R)
Lanes 3,5,7: S, C, E (F/MR)
Cut out ~250bp bands in lanes 2, 4, 6, and ~650bp bands in lanes 3, 5, 7. Gel extracted. Included extra QG step and eluted with 30ul EB.
Topo cloned/transformed using 4ul of gel extractions of Lpp29, OmpA66, and OmpA159, and 20ul Oneshot Top10 competent cells. Left in incubator at 3:00am.
7/19/06
Ran S, C, D(mut) on 120ml 1% agarose gel.
Lane 1: 1kb+
Lane 2, 3, 4: S, C, E(mut)
Cut out the ~350bp bands and gel extracted. Included extra QG step and eluted with 30ul EB.
Sent these gel extractions out for sequencing.
PT013: S(mut), StrepSCDF
PT014: S(mut), StrepSCDRS
PT015: C(mut), StrepSCDF
PT016: C(mut), SrepSCDRS
PT017: E(mut), StrepSCDF
PT018: E(mut), StrepSCDRS
Prepared eighteen new PCR reactions using PCR Supermix, NOT High Fidelity. (all primers at 5uM)
P1: 22ul PCR Supermix + 1ul StrepSCDF + 1ul StrepSCDMR + 1ul S, C, E (from 2ng/ul plasmid reconstitution 7/14/06)
G1: " + " + " + 1ul S, C, E (F/R) gel ex from 7/17/06
P2: 22ul PCR Supermix + 1ul StrepSCDRS + 1ul StrepSCDMF + 1ul S, C, E (recon)
G2: " + " + " + 1ul S, C, E (gel)
S, C, E (mut): 22ul PCR Supermix + 2ul S,C,E(F/MR) + 2ul S,C,E(R/MF) (gel ex 7/18/06)
Primer control 1: 22ul PCR Supermix + 1ul StrepSCDF + 1ul StrepSCDMR
Primer control 2: 22ul PCR Supermix + 1ul StrepSCDRS + 1ul StrepSCDMF
Primer control F/R: 22ul PCR Supermix + 1ul StrepSCDF + 1ul StrepSCDMRS
After 11 cycles (was supposed to be 10), I paused the machine, took out S, C, E (mut) and added 1ul StrepSCDF and 1ul StrepSCDRS to each, replaced them, and continued the PCR incubation.
Ran 1% 100ml agarose gel, 20 wells.
Lanes 1,20: 1kb+
Lanes 2,4,6: S,C,E(F/MR) (plasmid)
Lanes 3,5,7: S,C,E(R/MF) (plasmid)
Lanes 8,10,12: S,C,E(F/MR) (gel ex)
Lanes 9,11,13: S,C,E(R/MF) (gel ex)
Lanes 14-16: S,C,E(mut)
Lanes 17-19: Primer controls 1, 2, F/R
Made cuts around all visible bands in lanes 2-13, and left in 4dC overnight.
At around 7:00pm, I took out the Lpp29, OmpA66, and OmpA159 Topo cloning/transformation plates. I streaked and inoculated 3.3ml (33ul ampicillin, 5mg/ml) liquid cultures from one colony of each transformation. There weren't any carbinicillin plates, so I used a kanamycin plate; the Topo vector also carries kanamycin resistance.
7/20/06
Performed miniprep of Lpp29, OmpA66, OmpA159. Included PB step and eluted in 40ul water.
Prepared three digests. Master mix of 6ul water, 6ul Buffer 2, 1.2ul XbaI, 1.2ul PstI, 0.6ul BSA(100X). 15ul miniprep Lpp29,OmpA66,OmpA159 + 5ul master mix. Incubated at 37dC for 2h, 80dC for 20min.
Took out cut bands. 650bp from lanes 2,4,6: S1, C1, E1. 250bp from lanes 3,5,7: S2, C2, E2. Included extra QG step and eluted in 20ul EB.
Prepared three PCR reactions: 20ul Platinum PCR Supermix + 2ul S1,C1,E1(gel ex) + 2ul S2,C2,E2(gel ex). Paused after 10 cycles and added 1ul StrepSCDF and 1ul StrepSCDRS.
Ran 100ml 1% agarose gel, 130V, 45min, but the gel came out awful due to buffer that's been reused a few too many times.
Got sequences back for StrepW, H, D(mut). StrepW has a perfect sequence 1-444 from both M13F and M13R sequencing reactions. StrepH has a mutation at bp#344 T to C, which in the reading frame is a GCT to GCC, a silent mutation for Alanine. Strep D has a perfect sequence 1-409 from M13R reaction, and a perfect sequence 410-444 from M13F reaction.
Prepared StrepW,H,D(mut) cultures for midiprep in 100ml Terrific broth + 100ul kanamycin (50mg/ml). Also prepared new cultures of Lpp29, OmpA66, and OmpA159, 3ml LB + 30ml amipcillin (5mg/ml). Put in incubator at 6:30pm.
7/21/06
Removed liquid cultures at 10:30am.
Repeated PCR Reactions and digests from yesterday. Ran 1% 100ml agarose gel.
Lane 1,8: qkb+
Lanes 2-4: S,C,E(mut)
Lane 5: Lpp29 XbaI/PstI
Lane 6: OmpA66 XbaI/PstI
Lane 7: OmpA159 XbaI/PstI
Performed miniprep on Lpp29, OmpA66, OmpA159, including PB step and eluting in 30ul water. Sent off these minipreps for sequencing, 8ul for each reaction.
PT019: Lpp29, M13F(-21)
PT020: Lpp29, M13R
PT021: OmpA66, M13F(-21)
PT022: OmpA66, M13R
PT023: OmpA159, M13F(-21)
PT024: OmpA159, M13R
Performed midiprep on StrepW, StrepH, StrepD. Used N3 instead of P3, and skipped step 8, the 2nd centrifugation. Mixed more 70% ethanol (70ml of 95% ethanol + 25ml water).
7/24/06
Prepared liquid cultures (2ml LB + 20ul ampicillin 5mg/ml) of StrepW, StrepH, StrepD, and full OmpA, for glycerol stocks tomorrow.
Nanodropped StrepW, StrepH, StrepD midipreps. All 500-600 ng/ul.
Prepared three double digests. 0.3ul BSA(100X), 0.6ul XbaI, 0.6ul PstI, 2ul StrepW/H/D midiprep, 3ul Buffer2, 23.5ul water. Incubated at 37dC for 12h, 80dC for 20min.
Prepared three PCRs. 45ul PCR Platinum Supermix, 2ul StrepSCDF(5uM), 2ul StrepSCDRS(5uM), 1ul S/C/E(2ng/ul reconstitution from 7/14/06). PErformed PCR purification, eluting with 30ul water. Then prepared three digests: 0.35ul BSA, 0.7ul PstI, 3.5ul Buffer3, 30ul PCR-purified S/C/E. Incubated at 37dC for 8h, 80dC for 20min.
7/25/06
Made glycerol stocks of StrepW, StrepH, StrepD, full OmpA, with 1ml liquid culture + 666ul 50% glycerol. Froze in screwcap tubes at -80dC.
Used 1ul of the S,C,E PstI digests for three PCR ("before") reactions, plus 45ul Supermix, 2ul StrepSCDF(5uM), 2ul StrepSCDRS(5uM). Performed "PCR" purification on the remaining amount in the digest mixtures. Used 1ul of the purifications for three PCR ("after") reactions. Purified the S,C,E(before) and S,C,E(after) PCRs. Eluted in 30ul water, and prepared six digests with 30ul purified sample plus 5ul from a master mix of 7x(0.35ul 100X BSA, 0.45ul water, 0.7ul PstI, 3.5ul Buffer3). Incubated at 37dC for 8h, 80dC for 20min.
Topo cloned/transformed S,C,E(F/R) gel extractions for 7/17/06, in preparation for arrival of QuikChange Multi site-directed mutagenesis kit. Left plates in incubator around 6:00pm.
Got sequences back for Lpp29, OmpA66, and OmpA159. Analyzed and confirmed correct sequences for Lpp29 and OmpA66, and one silent mutation in OmpA159. Prepared 100ml liquid cultures + 100ul kanamycin (50mg/ml). Used LB for OmpA66 and OmpA159, and 92ml Terrific broth + 8ml LB for Lpp29. Left in incubator around 7:00pm.
Ran gel of StrepW,H,D XbaI/PstI digests, and purified S,C,E PCR/digests. I tried loading 20ul of the purified S,C,E PCR/digests + 2ul 10X loading dye, but for some reason they all diffused out of the wells.
Lanes 1,5: 1kb+
Lanes 2-4: purified S,C,E PCR/digests
Lanes 6-8: StrepW,H,D XbaI/PstI digests
Cut out ~400bp bands from lanes 6-8. Left in freezer.
7/26/06
Streaked and inoculated 3ml cultures from yesterday's Topo cloning/transformation of S,C,E(F/R).
Performed midiprep of Lpp29, OmpA66, OmpA159. I spilled the final OmpA159 elution, but I was able to recover ~100ul.
Purified the S,C,E(after) digests from last night.
Prepared six PCRs. 45ul Supermix + 2ul StrepSCDF + 2ul StrepSCDRS + 1ul S,C,E(before) digests or 1ul S,C,E(after) purified digests.
Prepared three more XbaI/PstI digests of StrepW, H, D midiprep. Same as yesterday, but used 3ul midiprep and 22.5ul water, and incubated at 37dC for 2h.
Ran a 1% 100ml agarose gel, 130V, 45min.
Lanes 1,5,9: 1kb+
Lanes 2-4: S,C,E(before) digests
Lanes 6-8: S,C,E(after) digests
Lanes 10-12: StrepW,H,D XbaI/PstI digests
As a reminder, the S,C,E(before) digests had the following history: PCR, purified, PstI digest, not purified and 1ul used in PCR, purified, PstI digest. S,C,E(after) digests had the following history: PCR, purified, PstI digest, purified and 1ul used in PCR, purified, PstI digest, and purified once more (in preparation for the PCRs detailed above). I added 4ul loading dye to each sample before loading into the gel. The S,C,E(after) purified digests floated out of their wells.
Cut out the 350bp band from lanes 2-4 and the 400bp band from lanes 10-12.
Prepared six double digests. R0010, E0241, and E7104 were minipreps provided by David Ramos and Zhipeng Sun. Incubated at 37dC for 12h, 80dC for 20min.
25ul DNA + 3ul buffer + 0.6ul enzyme 1 + 0.6ul enzyme 2 + 0.3ul BSA
- R0010 miniprep, EcoRI/PstI, Buffer 3
- E0241 miniprep, EcoRI/SpeI, Buffer 2
- E0741 miniprep, XbaI/PstI, Buffer 2
- 2ul Lpp29 midiprep + 23ul water, EcoRI/SpeI, Buffer 2
- 3ul OmpA66 midiprep + 22ul water, XbaI/PstI, Buffer 2
- 3ul OmpA159 midiprep + 22ul water, XbaI/PstI, Buffer 2
7/27/06
Performed minipreps of S/C/E(F/R).
Ran digests on 1% agarose gel.
Lanes 1,8: 1kb+
Lane 2: R0010 E/P
Lane 3: E0241 E/S
Lane 4: E0741 X/P
Lane 5: Lpp29 E/S
Lane 6: OmpA66 X/P
Lane 7: OmpA150 X/P
Cut out 2kb band from lanes 2-4 (lane 2 is actually two bands very close to each other; I cut out the lower one). Cut out 120, 100, 400bp bands from lanes 5,6,7. Gel purified, included extra QG step and eluted with 30ul EB.
Performed five ligation/transformations, using 24ul aliquots of Top10 OneShot. After addition of ligation mixture, I incubated them on ice for 1h, as opposed to 30min normally; and during initial growth, I incubated them on a shaker at 37dC for 1.5h, as opposed to 1h normally.
- 2ul R0010 E/P + 4ul Lpp29 E/S + 4ul OmpA66 X/P
- 2ul R0010 E/P + 4ul Lpp29 E/S + 4ul OmpA150 X/P
- 2.5ul E0241 E/S + 7.5ul Lpp29 E/S
- 2.5ul E0741 X/P + 7.5ul OmpA66 X/P
- 2.5ul E0741 X/P + 7.5ul OmpA150 X/P
7/28/06
Streaked and inoculated 3ml liquid cultures + 30ul amp (5mg/ml) from ligation/transformations yesterday. Left in incu-shaker at 11:00am.
Received primers for QuikChange mutagenesis. Reconstituted StrepSCDMF2 in 939ul EB for 20uM, then made another dilution of 25ul 20uM + 75ul water for 5uM.
Diluted S,C,E(F/R) minipreps. According to the Nanodrop, S was 220ng/ul, C 160, E 220. So I mixed 3ul S with 9ul water for 1in4 dilution, 3 C with 6 for 1in3, 3E with 9 for 1in4. This was to obtain a concentration of ~50ng/ul.
Prepared mixtures for Quikchange multi site-directed mutagenesis. 2.5ul buffer, 1ul dNTP, 1ul enzyme, 15.34ul water, 1ul S/C/E, 4.16ul StrepSCDMF2 primer. I used 4.16ul primer b/c the protocol suggested 100ng per primer if using 1-3 primers., and I was sure that the primer would bind to two homologous sites and act like two primers. I incubated the mixtures according to the protocol: 95dC for 1min, 30x(95dC for 1min, 55dC for 1 min, 65dC for 10 min), and cool down. A time of 10min was chosen for the 65dC step b/c the protocol called for 2min/kb of plasmid length. The vectors are 3.9kb pCR 2.1-Topo vector + ~900bp insert = ~5kb. Then DpnI digestion according to protocol.
I prepared three transformations with the XL10-Gold ultracompetent cells according to the protocol, except that I used 500ul SOC media instead of NZY+ broth. I also prepared three transformations of Top10 OneShot cells (20ul aliquots, 1.5ul DNA, 250ul SOC) in parallel. Plated 100ul of XL10-Gold transformations and total volume of Top10 transformations; left at 37dC at 10:30pm. I refrigerated the remaining volume of XL10-Gold transformations at 4dC.
Performed minipreps of Lpp29, OmpA66, OmpA159, Lpp29-OmpA66, Lpp29-OmpA159, in pSB1A2. Prepared five 25ul digests. Master mix was 6x(2.5ul Buffer2, 1.25ul water, 0.5ul EcoRI, 0.5ul SpeI, 0.25ul BSA(100X)). 20ul miniprep + 5ul master mix. Incubated at 37dC for 10h, 80dC for 20min.
7/29/06
Colonies appeared on all three transformation plates of mutated S,C,E in XL10-Gold, and on plates of S and C in Top10, but not on the plate of E in Top10. Baffling. Did I forget to add DNA to that tube? Speaking of the DNA, I had meant to freeze the mutagenesis mixture, but I left it out at room temperature overnight for ~15h.
Streaked and inoculated 3ml cultures from a single colony on S,C,E (XL10) and S,C(Top10) transformation plates. Left in incu-shaker at 3:00pm.
7/30/06
Took liquid cultures out at 9:00am, 18h in incu-shaker. Performed minipreps, including PB step and eluted with 40ul water.
Prepared five 25ul double digests. 20ul miniprep of (S,C,E(XL);S,C(Top)) + 5ul of master mix, which was 6x(2.5ul Buffer3, 1.25ul water, 0.5ul XbaI, 0.5ul PstI, 0.25ul BSA(100X)). I usually use Buffer 2 in which XbaI is 100% efficient and PstI is 75% efficient, but because I'm testing for the mutation of a PstI site, I decided to use Buffer3 in which XbaI is 75% efficient and PstI is 100% efficient. Incubated at 37dC for 6h, 80dC for 20min.
Ran 1% 100ml gel of digests from today and 7/28/06.
Lanes 1,7: 1kb+
Lane 2: Lpp29 E/S
Lane 3: OmpA66 E/S
Lane 4: OmpA159 E/S
Lane 5: Lpp29-OmpA66 E/S
Lane 6: Lpp29-OmpA159 E/S
Lane 8: SCD-NM mut (XL10 transf)
Lane 9: C2 mut (XL10 transf)
Lane 10: E2 mut (XL10 transf)
Lane 11: SCD-NM mut (Top10 transf)
Lane 12: C2 mut (Top10 transf)
7/31/06
Sent out stuff for sequencing.
- PT025: Lpp29 in pSB1A2, VF2
- PT026: OmpA66 in pSB1A2, VF2
- PT027: OmpA159 in pSB1A2, VF2
- PT028: Lpp29-OmpA66 in pSB1A2, VF2
- PT029: Lpp29-OmpA159 in pSB1A2, VF2
- PT030: SCD-NM mut in Topo from Top10, M13F
- PT031: SCD-NM mut in Topo from Top10, M13R
- PT032: SCD-NM mut in Topo from XL10, M13F
- PT033: SCD-NM mut in Topo from XL10, M13R
Prepared eight 3ml liquid cultures. Aliquotted from mix of 26ml LB + 260ul amp (5mg/ml). Left in incu-shaker at 4:30pm.
- Lpp29, OmpA159, Lpp29-OmpA66 from 7/28/06 streaks.
- R0010 from 6/12/06 transformation; streaked onto a carb plate.
- C.1, C.2, E.1, E.2: two colonies each from C2 and E2(non-mut)transformations 7/25/06; streaked onto a carb plate.
Plans: Digest Lpp29 S/P, OmpA159 X/P for suffix insertion. Digest Lpp29-OmpA66 S/P for suffix insertion of StrepW,H,D. Digest R0010 S/P for eventual suffix insertion and expression of construct. Digest C.1, C.2, E.1, E.2 X/S for insert check.
8/1/06
Performed minipreps of the liquid cultures yesterday. I spilled a drop of Lpp29 culture while resuspending in P1. I forgot about the streak plate until 6:00pm, so they were incubated for ~25h; there are visible satellite colonies around the streaks.
Prepared eight 25ul digests. 20ul miniprep + 2.5ul Buffer2 + 1.25ul water + 0.5ul enzyme1 + 0.5ul enzyme2 + 0.25ul BSA(100X).
- Lpp29, Lpp29-OmpA66, R0010 S/P as vectors for suffix insertion
- C.1, C.2, E.1, E.2 X/S for insert check
- OmpA159 X/P as insert for suffix insertion
Ran a 1% 100ml agarose gel, 130V, 45min.
Lanes 1,6: 1kb+
Lanes 2-5: C.1, C.2, E.1, E.2 X/S
Lane 7: Lpp29 S/P
Lane 8: OmpA159 X/P
Lane 9: Lpp29-OmpA66: S/P
Lane 10: R0010 S/P
Cut out ~2kb bands from Lanes 7,9,10, and ~400bp band from lane 8. Performed gel purification, including extra QG step and eluting with 30ul EB.
Performed alkaline phosphatase treatment. Added 3.5 10X buffer and 1ul alkaline phosphatase. Put in 37dC incu-shaker for 1h, then 65dC for 10min in PCR machine.
Confirmed sequences for Lpp29, OmpA66, OmpA159, Lpp29-OmpA66 in pSB1A2 vectors. Did not get a correct band size for Lpp26-OmpA159 digest check, and sequence is not correct.
Prepared 26ml LB + 260ul amp (5mg/ml) liquid cultures of Lpp29, OmpA66, OmpA159, Lpp29-OmpA66 in pSB1A2, for glycerol stock and midiprep.
Performed seven ligation/transformations, with 2.5ul vector + 7.5ul insert, and 15ul aliquots of Top10.
- Lpp29 S/P + OmpA159 X/P
- Lpp29-OmpA66 S/P + StrepW,H,D X/P
- E0741 X/P + StrepW,H,D X/P
Dissolved BBa_B0032 from well 3K on iGEM plate DNA-1 with 15ul water, then transformed 1ul into 15ul aliquot of Top10.
For ligation, I used 2ul of a 1:80 dilution of NEB T4 DNA ligase because the DNA ligase in the kit ran out.
During the transformations, I left the transforming cells on ice for a little over 1h before heat shock, I didn't leave on ice for a full 2min after heat shock, and I added 200ul SOC.
Prepared twelve 3ml LB + 3ul cultures. Made master mix of 40ul LB + 40ul kanamycin (50mg/ml).
- SCD-NM mutagenized from XL10, from Top10
- C3-C7, streaked and inoculated from colonies on C(F/R) Topo cloning/transformation 7/25/06
- E3-E7, streaked and inoculated from colonies on E(F/R) Topo cloning/transformation 7/25/06
8/2/06
Made glycerol stocks of Lpp29, OmpA66, OmpA159, Lpp29-OmpA66. I decided to hold off on a midiprep.
Performed minipreps of SCD-NM mut from XL10, SCD-NM mut from Top10, C3,6,7, and E3,6,7. Prepared six 25ul X/S digests of C3,6,7 and E3,6,7; incubated at 37dC for 2h, 80dC for 20min.
Ligation/transformations yielded very few colonies. Used inoculation needle to pick a colony, streak, and inoculate a 3ml LB + 30ul amp(5mg/ml) liquid culture, then attempted a colony PCR by running the needle over the streak again and dipping into a mix of 16ul PCR Supermix + 2ul VF2(2uM) + 2ul VR (2uM).
- Lpp29-OmpA66-StrepW,H,D
- StrepW,H,D in pSB1A2
- Lpp29-OmpA159
B0032 transformation yielded many colonies. Made a streak and a 3ml LB + 3ul amp(5mg/ml) liquid culture.
Received constructs 481 (Lpp1-9-OmpA46-66-Bla fusion) and 488 (Lpp1-9-OmpA46-145-Bla fusion) from George Georgiou, in tubes on solid agar. Made a streak and a 3ml LB + 3ul amp(5mg/ml) liquid culture, and attempted a colony PCR by running the needle over the streak again and dipping into a mix of 45ul PCR Supermix + 2ul LppFS(5uM) + 2ul OmpA66R/OmpA159R(5uM).
Received BBF primer, which anneals only to the upstream BioBricks site. Reconstituted in 393ul EB for 100uM, then diluted 5uM of that with 95ul EB for 5uM. Prepared six PCR reactions: 2x ( 45ul PCR Supermix + 2ul BBF(5uM) + 2ul StrepSCDRS(5uM) + 1ul S,C,E(F/R no mut 7/17/06) ). One set for Topo cloning, one set for PstI selection.
Ran a 100ml 1% agarose gel, 130V, 40min.
Lanes 1,9: 1kb+
Lanes 2-4: StrepW,H,D in pSB1A2, PCR
Lanes 5-7: Lpp29-OmpA66-StrepW,H,D in pSB1A2, PCR
Lane 8: Lpp29-OmpA159
Ran a 1.2% e-gel. Took 10ul of samples and mixed with 10ul water.
Lane 1: 1kb+
Lane 2-4: C3,6,7 XS digest
Lanes 5-7: E3,6,7 XS digest
Lanes 9-10: S,C,E(no mut) PCR
Lane 11: Construct 481 PCR
Lanes 12: Construct 488 PCR
8/3/06
Made 10ul colony PCRs, 8ul PCR Supermix + 1ul forward primer(2uM) + 1ul reverse primer(2uM), and plated streaks from colonies of ligation/transformations. Ran 1.2% e-gels.
Lane 1: 1kb+
Lanes 2-3: LO66W #2,3 (VF2/VR)
Lanes 4-7: LO66 #2,3,4,5
Lanes 8-10: LO66D #2,3,4
Lanes 11-12: LO159 #2,3
Lane 1: 1kb+
Lanes 2-3: C2 #4,5 (M13F,M13R)
Lanes 4: W #2 (VF2/VR)
Lanes 5-9: H #2,3,4,5,6
Lanes 10-12: D #1,2,3
The 481 streak grew, while the 488 streak did not. Both liquid cultures grew, so I used an inoculation needle and plated a streak from the 488 liquid culture.
Prepared 3ml LB + 30ul amp(5mg/ml) cultures from LO66W(#3) streak for sequencing, and also SCD-NM, C2, E2 in pET vector, streaked on 7/15/06, for QuikChange mutagenesis. Left in incu-shaker around 12:30pm.
Performed minipreps of B0032, 481, 488, W,H,D, LO159, LO66W cultures.
Sent miniprepped DNA for sequencing.
- PT034: B0032, VF2
- PT035: LO159, VF2
- PT036: W in pSB1A2, VF2
- PT037: W in pSB1A2, VR
- PT038: LO66W(#3), VF2
- PT039: LO66W(#3), VR
- PT040: D in pSB1A2, VF2
- PT041: D in pSB1A2, VR
- PT042: SCD-NM(mut) from Top10, M13F
- PT043: SCD-NM(mut) from Top10, M13R
- PT044: SCD-NM(mut) from XL10, M13F
- PT045: SCD-NM(mut) from XL10, M13R
Prepared colony PCRs/streaks, and ran an e-gel.
Lane 1: 1kb+
Lanes 2-4: LO66H #6,7,8
Lanes 5-8: H #7,8,9,10
Lanes 9-12: D #4,5,6,7
Lane 1: 1kb+
Lane 2-12: C #8-18 (C#16 did not have anything in its PCR tube. Curious.)
Prepared 100ml LB cultures + 100ul amp(50mg/ml) of Lpp29, OmpA66, OmpA159, Lpp29-OmpA66, for midiprep tomorrow. Left in incu-shaker around 5:30pm.
Prepared 3ml LB cultures + 31ul amp(5mg/ml) of SCD-NM (original Topo clone/transf), C2 (from colony #8's streak), E2 (from colony #7's streak), in preparation for miniprep and Quikchange mutagenesis.
Performed miniprep of LO66W (#3) and of SCD-NM, C2, E2 in their original pET vectors, for following mutagenesis.
Prepared Quikchange mutagenesis reactions. 2.5ul buffer, 1ul dNTP, 1ul enzyme, 15.34ul water, 1ul S/E (50ng/ul), 4.16ul StrepSCDMF2 primer (5uM). I used 14.34ul water and 2ul for C(25ng/ul). I realized right before adding enzyme that I needed to add Quik solution because the template is 5.5kb pET22b(+) vector + 800bp insert = ~6.5kb. I added 0.75ul Quiksolution, and then finally the enzyme. I incubated according to protocol, with the 65dC step timed for 13min.
8/4/06
Finished Quikchange mutagenesis of SCD-NM, C2, E2 in pET22 by treating with Dpn1, then transformed into XL10 cells by protocol, except for using SOC media instead of NZY+ broth. Also did a parallel transformation with 1.5ul of Dpn1-treated DNA in 20ul aliquots of Top10 cells. Left plates in incubation around 2:30pm.
Performed midipreps of Lpp29, OmpA66, OmpA159, and Lpp29-OmpA66 in pSB1A2.
Performed minipreps of SCD-NM, C2(#8), E2(#7) in TOPO vector, not mutated.
Rec'd primers Strep13F and Strep13RS for pTSA13 clone from Dr. Sano. Reconstituted in EB to 100uM, then made a 5uM dilution. I reconstituted the pTSA13 DNA in 50ul water for ~2ng/ul, and transferred to a new tube (the tube it arrived in was cracked). Prepared a PCR of 45ul PCR Supermix + 2ul Strep13F(5uM) + 2ul Strep13RS(5uM) + 1ul pTSA13 DNA.
Made 2uM dilutions of M13F(-20) and M13R primers that come with the Topo-TA cloning kit. Prepared three PCR reactions: 15ul PCR Supermix + 2ul M13F(-20)(2uM) + 2ul M13R(2uM) + 1ul SCD-NM,C2,E2 miniprep.
Ran a 1% 100ml agarose gel, 130V, 45min (Mingming stopped and imaged the gel).
Lane 1: 1kb+
Lanes 2-3: pTSA13 PCR, split up into two wells
Lanes 4-6: S, C, E in Topo vector, M13F/R PCR
Hetmann cut out the ~300bp bands from lanes 2-3 and gel extracted.
Prepared streaks and 3ml LB + 3ul amp(50mg/ml) cultures of SCD-NM, C2, E2(mut in pET22) from XL10 and Top10 transformations.
8/5/06
Mingming took out streak and cultures and performed minipreps.
8/9/06

0.5x TBE
| Lane | Contents | Loading Buffer |
| 0 | 1kb ladder (10 μL) | - |
| 1 | OmpA159 (28 μL) | 3 μL |
| 2 | R0010 (28 μL) | 3 μL |
| 3 | B0032 (28 μL) | 3 μL |
| 4 | Lpp29 (33 μL) | 4 μL |
| 5 | OmpA159+Lpp29 (33 μL) | 4 μL |
| 6 | S #1 (28 μL) | 6 μL |
| 7 | S #2 (28 μL) | 6 μL |
| 8 | E #1 (28 μL) | 6 μL |
| 9 | E #2 (28 μL) | 6 μL |
| 10 | C #1 (28 μL) | 6 μL |
| 11 | C #2 (28 μL) | 6 μL |
| 12 | 481 #1 (28 μL) | 6 μL |
| 13 | 481 #2 (28 μL) | 6 μL |
| 14 | 488 #1 (28 μL) | 6 μL |
| 15 | 488 #2 (28 μL) | 6 μL |
8/16/06
Prepared colony PCRs, 8ul PCR Supermix + 2ul M13F(2uM) + 2ul M13R(2uM), and kan plate streaks from three colonies each on transformation plates for C(mut), E(mut), pTSA13 from last week.
Prepared 3ml LB + 30ul amp(5mg/ml) liquid cultures and streaks of R0010 (from 6/12/06 transformation) and of B0032(from 8/2/06 streak), and Lpp29-OmpA159, StrepH, Lpp29-OmpA66-StrepD in pSB1A2 and S(mut) in pCR-2.1 Topo (from last week's transformations).
8/17/06
Performed minipreps of liquid cultures. Prepared 25ul digests: 20ul miniprep, 2.5ul Buffer2, 1.25 ul water, 0.5ul enzyme1, 0.5ul enzyme2, 0.5ul BSA(100X).
- R0010, E/S
- B0032, E/X
- Lpp29-OmpA159, S/P
- StrepH, X/P
- Lpp29-OmpA66-StrepD, X/P
- S(mut), X/P
Treated B0032 (E/X) and Lpp29-OmpA159 (S/P) with 1ul CIP + 3ul dephosphorylation buffer, for 1h at 37dC.
Ran 1% agarose gel.
Lane 1: 1kb+
Lane 2: R0010, E/S
Lane 3: S(mut), X/P
Lane 4: Lpp29-OmpA66-StrepD, X/P
Lane 5: StrepH, X/P
Lane 6: B0032, E/X (CIPed)
Lane 7: Lpp29-OmpA159, S/P (CIPed)
Cut out the following bands: 250bp R0010, 850bp S(mut), 600bp L-O66-D, 2kb B0032, 2.5kb L-O159. Left gel slices in freezer.
Ran 1.2% e-gel of colony PCRs.
Lane 1: 1kb+ ladder
Lanes 2-4: C1,2,3
Lanes 5-7: E1,2,3
Lanes 8-10: pTSA13 1,2,3
Lanes 11,12: water
Sent minipreps for sequencing.
- PT046: Lpp29-OmpA159 in pSB1A2, VF2
- PT047: Lpp29-OmpA159 in pSB1A2, VR
- PT048: StrepH in pSB1A2, VF2
- PT049: StrepH in pSB1A2, VR
- PT050: Lpp29-OmpA66-StrepD in pSB1A2, VF2
- PT051: Lpp29-OmpA66-StrepD in pSB1A2, VR
- PT052: S mut in pCR-2.1 Topo, M13F(-21) (7ul)
- PT053: S mut in pCR-2.1 Topo, M13R (6ul)
Prepared 3ml LB + 3ul amp(5mg/ml) liquid cultures of E(#1) and pTSA13(#1). Prepared 11 colony PCRs and kan plate streaks from colonies on C transformation plate. Ran another 1.2% e-gel, all bands were incorrect at around 200bp. Repeated the process with another 11 colony PCRs and streaks. Again, an e-gel showed all incorrect bands. UGH.
8/18/06
Performed minipreps of E and pTSA13, sent out for sequencing.
- PT054: E(mut), M13F
- PT055: E(mut), M13R
- PT056: pTSA13(mut), M13R
- PT057: pTSA13(mut), M13F
Performed gel extractions of bands cut out yesterday, and also of old gel slices of StrepW, H, D.
Performed ligation/transformations with 15ul aliquots of Top10 cells. Poured my own amp plates.
- R0010 E/S + B0032 E/X
- Lpp29-OmpA66 S/P + StrepW X/P
- Lpp29-OmpA66 S/P + StrepH X/P
- Lpp29-OmpA66 S/P + SCDNM X/P
- Lpp29-OmpA159 S/P + StrepW X/P
- Lpp29-OmpA159 S/P + StrepH X/P
- Lpp29-OmpA159 S/P + StrepD X/P
- Lpp29-OmpA159 S/P + SCDNM X/P
8/19/06
Prepared colony PCRs, 8ul PCR Supermix + 1ul VF2(2uM) + 1ul VR(2uM), and amp plate streaks of yesterday's ligation/transformations.
8/20/06
Ran 1.2% e-gel of colony PCRs.
Lane 1: 1kb+
Lane 2: R0010-B0032
Lane 3: Lpp29-OmpA66-StrepW
Lane 4: Lpp29-OmpA66-StrepH
Lane 5: Lpp29-OmpA66-SCDNM
Lane 6: Lpp29-OmpA159-StrepW
Lane 7: Lpp29-OmpA159-StrepH
Lane 8: Lpp29-OmpA159-StrepD
Lane 9: Lpp29-OmpA159-SCDNM
Lanes 10-12: water
Prepared 3ml + 30ul amp(5mg/ml) liquid cultures of Lpp29-OmpA66-StrepW, Lpp29-OmpA66-StrepH, Lpp29-OmpA159-StrepW, Lpp29-OmpA159-StrepD, for miniprep/sequencing tomorrow.
Prepared new 10ul colony PCRs/amp plate streaks from R0010-B0032 (3 colonies), Lpp29-OmpA66-SCDNM (3), Lpp29-OmpA159-StrepH (2), Lpp29-OmpA159-SCDNM (3).
8/21/06
Ran 1.2% e-gel of colony PCRs.
Lane 1: 1kb+
Lanes 2-4: R0010-B0032 #1-3
Lanes 5-7: LO66S #1-3
Lanes 8-10: LO159S #1-3
Lanes 11-12: LO159H #1,2
Prepared 3ml LB + 30ul amp(5mg/ml) liquid cultures of R0010-B0032, LO66S, LO159S, LO159H #1, for miniprep/sequencing.
Prepared 30ml LB + 30ul amp(50mg/ml) liquid culture Lpp29-OmpA159, for midiprep.
Performed minipreps of liquid cultures from yesterday. Eluted in 40ul water. Sent out for sequencing.
- PT058: LO66W, VF2
- PT059: LO66W, VR
- PT060: LO66H, VF2
- PT061: LO66H, VR
- PT062: LO159W, VF2
- PT063: LO159H, VR
- PT064: LO159D, VF2
- PT065: LO159D, VR
8/22/06
Performed minipreps of 3ml liquid cultures from yesterday. I usually centrifuge 1.5ml twice, and when I was pipetting in liquid culture for the 2nd centrifugation, I accidentally pipetted R0010-B0032 culture into the tube with the 1st LO159H pellet. I poured the R0010-B0032 culture back into its culture tube, and then pipetted into the correct tube for centrifugation. I shook out the remainder of R0010-B0032 liquid from the LO159H tube. I continued the miniprep normally, and eluted in 40ul. This means I have definite, but hopefully minor, cross-contamination of these R0010-B0032 and Lpp29-OmpA159-StrepH minipreps.
Sent out for sequencing.
- PT066: LO66S, VF2
- PT067: LO66S, VR
- PT068: LO159H, VF2
- PT069: LO159H, VR
- PT070: LO159S, VF2
- PT071: LO159S, VR
- PT072: R0010-B0032, VF2
Performed midiprep of Lpp29-OmpA159, along with cyano's J04500.
Prepared 25ul digests: 20ul DNA, 2.5ul Buffer2, 1.25 ul water, 0.5ul enzyme1, 0.5ul enzyme2, 0.5ul BSA(100X), from minipreps today and yesterday, today's LO159 and J04500 midipreps, and a previous LO66 midiprep.
XbaI/PstI digests.
- LO66 (4ul midiprep 275 ng/ul, 16ul water)
- LO159 (6.67ul midiprep 150ng/ul, 13.33ul water)
- LO66W
- LO66H
- LO66S
- LO159W
- LO159H
- LO159D
- LO159S
SpeI/PstI digests.
- R0010-B0032
- J04500 (midiprep I included today for cyano group: 4ul midiprep 240 ng/ul, 16ul water)
8/23/06
Ran 1% 100ml agarose gel of digests.
Lane 1: 1kb+
Lane 2: R0010-B0032
Lane 3: J04500
Lane 4: LO66
Lane 5: LO159
Lane 6-8: LO66W, H, S
Lane 9-12: LO159W, H, D, S
Seems like R0010-B0032 was not digested. Cut out J04500 band, and the lower band from the rest of the lanes. Performed gel extractions. Eluted with 30ul water.
Performed eight ligation/transformations, using 14ul aliquots of Top10F' competent cells.
- J04500 S/P + LO66 X/P
- J04500 S/P + LO159 X/P
- J04500 S/P + LO66W X/P
- J04500 S/P + LO66H X/P
- J04500 S/P + LO66S X/P
- J04500 S/P + LO159W X/P
- J04500 S/P + LO159H X/P
- J04500 S/P + LO159S X/P
8/24/06
Took plates out of incubation, and prepared 10ul colony PCRs and carb plate streaks from two colonies of each transformation.
Ran PCRs on 1% 80ml agarose gel, 20 wells.
Lane 1,18: 1kb+
Lanes 2,3: JLO66 1,2
Lanes 4,5: JLO66W 1,2
Lanes 6,7: JLO66H 1,2
Lanes 8,9: JLO66S 1,2
Lanes 10,11: JLO159 1,2
Lanes 12,13: JLO159W 1,2
Lanes 14,15: JLO159H 1,2
Lanes 16,17: JLO159S 1,2
Prepared 3ml LB + 0.85ul amp(50mg/ml) liquid cultures of all #1 colonies. I had accidentally made a mix of 35ml + 30ul amp(50mg/ml).
Mingming performed a miniprep of R0010-B0032.
8/25/06
Performed minipreps of liquid cultures and sent out for sequencing.
- PT073: JLO66 VF2
- PT074: JLO66 VR
- PT075: JLO66W VF2
- PT076: JLO66W VR
- PT077: JLO66H VF2
- PT078: JLO66H VR
- PT079: JLO159 VF2
- PT080: JLO159 VR
- PT081: JLO66S VF2
- PT082: JLO66S StrepSCDF
- PT083: JLO66S VR
- PT084: JLO159W VF2
- PT085: JLO159W OmpA159R
- PT086: JLO159W VR
- PT087: JLO159H VF2
- PT088: JLO159H OmpA159R
- PT089: JLO159H VR
- PT090: JLO159S VF2
- PT091: JLO159S OmpA159R
- PT092: JLO159S StrepSCDF
- PT093: JLO159S VR
Prepared a 25ul digest of R0010-B0032.
8/28/06
I realized that I left the streaks of my precious constructs in the incubator over the weekend, so they overgrew.
Prepared 2ml LB amp cultures of JLO66, JLO66W, JLO66S, JLO159, JLO159W, JLO159S. Also prepared 2ml LB amp + 1mM IPTG liquid cultures. Also made new carb streaks.
8/29/06
Of the IPTG cultures, JLO66W and JLO159W and JLO159 grew well, JLO66 and JLO66S grew less well, and JLO159S did not grow at all (toothpick stick on the side, though?).
Of the streaks, the JLO66S streak did not grow well, and LO159S barely grew at all.
All non-IPTG cultures grew well. Left in refrigeration.
8/30/06
JLO66, JLO66W, JLO66S, JLO159, JLO159W, JLO159S
Made 1:20 dilutions of cultures in 1ml LB amp. Incubated for 1h. Added 2ul 0.5M IPTG, incubated for 2h. Took 200ul samples for experiment, and another set 200ul samples for negatives. Pelleted, resuspended in 100ul PBS. Added 2ul 100uM "S0" (streptavidin aptamer with rhodamine tag). Incubated on rotisserie at room temperature for 1.5h. Pelleted, and washed 3x with PBS. Then resuspended in 200ul PBS and loaded into 96well blk/clrbtm plate.
Performed minipreps of E(mut) and pTSA13. Prepared X/P digests of E(mut) and pTSA13, and an E/S digest of pTSA13 because the sequencing results show a mutation of the PstI site.
CIPed the R0010-B0032 digest.
8/31/06
Performed new colony VF2/VR PCRs and streaks of the JLO constructs. Ran a gel of those and of the digests.
Lane 1: 10ul 1kb+
Lane 2: R0010-B0032 S/P
Lane 3: pTSA13 E/S
Lane 4: pTSA13 X/P
Lane 5: E(mut) X/P
Lanes 6-9: LO66, LO66W, LO66H, LO66S colony PCRs
Lanes 10-13: LO159, LO159W, LO159H, LO159S colony PCRs
Lane 14: 5ul 1kb+
Prepared 2ml LB amp cultures for glycerol stock tomorrow, picking from plate streaks.
- R0010 in pSB1A2 in Top10, 8/16/06
- B0032 8/16/06
- R0010-B0032 8/20/06
- Lpp29-OmpA159, 8/12/06
- LO66W 8/19/06
- LO66H 8/19/06
- LO66D 8/12/06
- LO66S 8/20/06
- LO159W 8/19/06
- LO159H 8/20/06
- LO159D 8/19/06
- LO159S 8/20/06
- SCD-NM in pCR 2.1 Topo 8/12/06
- E2 in pCR 2.1 Topo 8/16/06
- pTSA13 in pCR 2.1 Topo 8/16/06
9/1/06
Made the glycerol stocks, 900ul culture + 600ul 50% glycerol. Stored in -80dC freezer.
Prepared another set of colony PCRs from the streaks. Ran on a 70ml, 20well 1% agrose gel (maybe 10ml of the dissolved agarose solidified inside the bottle, so maybe only 60ml gel).
Lane 1: 10ul 1kb+
Lanes 2-5: LO66, LO66W, LO66H, LO66S
Lanes 6-9: LO159, LO159W, LO159H, LO159S
Lane 10: 5ul 1kb+