BMCB625:DNA Gyrase
Discussion/Homework Questions
Mahta
Q1. It takes 20x higher concentration of gyrase to relax the DNA in conditions lacking ATP (passive mode) as opposed to the negative 'supercoiling' ATP-dependent mode. Is their choice of DNA without a high affinity gyrase binding site valid for a more "in vivo" representation of the two different modes? Why use DNA lacking a strong gyrase recognition site?
Q2. It's stated that, "our single-molecule experiments reveal that the continued ATP hydrolysis may alternatively derive from a combination of two modes of catalysis, one that introduces (-) supercoils using ATP and a second that removes them without the use of ATP..." Is this describing a state of equilibrium? Is the ATP-independent activity more robust in the presence of ATP? (had a hard time wording this question..)
Larry
Jon
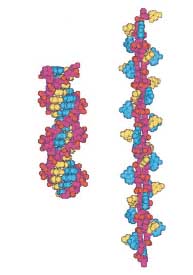
Q: How might the relaxation of + supercoils with the addition of force differ from that of - supercoiling relaxation with the additional observation of pauling like structures exist (see below right) when positive supercoiled DNA is streched by 3 pN of force.
- Allemand JF, Bensimon D, Lavery R, and Croquette V. Stretched and overwound DNA forms a Pauling-like structure with exposed bases. Proc Natl Acad Sci U S A. 1998 Nov 24;95(24):14152-7. DOI:10.1073/pnas.95.24.14152 |
Jeremy
Q1. Why and how might an antimicrobial target DNA gyrase?
Part 1. Fortunately, bacterial gyrases differ significantly from our gyrases, making them an attractive antimicrobial target.
Part 2. Potential targets: ATP binding or hydrolysis, DNA cutting, binding or wrapping.
Chris
Discussion point: Historically, DNA circles have been found to be underwound, that is “their linking numbers are less than those of their corresponding relaxed circles. This phenomena has been established by observing the effect of ethidium ion (from EtBr) affecting the rate of sedimentation (Voet & Voet 3/e).” This has been shown experimentally by plotting the Buoyant density as a function of EtBr concentration (in ug/uL).
How does today’s paper support this biophysical finding?