BME100 f2013:W900 Group2 L4
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | |
|
OUR TEAMLAB 1 WRITE-UPInitial Machine TestingThe Original Design
Experimenting With the Connections When part 3 was unplugged from part 6, the machine's screen turned off. This would imply that this is the power source for the screen readout. The white wire connecting part 6 to part 2 was the temperature sensor, so when unplugged, the machine became unable to read temperature, and the temperature changed from 0 degrees Celsius to -63.9 degrees Celsius.
The first test of the OpenPCR machine on October 23, 2013 at 10:07 AM. The machine was in working order and performed to the expectations given by our instructors. The machine performed 22 cycles of DNA replication in one hour and seven minutes. There was no malfunction in heat cycles or starting the machine.
ProtocolsThermal Cycler Program
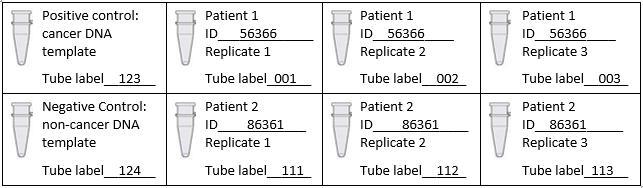
DNA/ primer mix
Research and DevelopmentPCR - The Underlying Technology
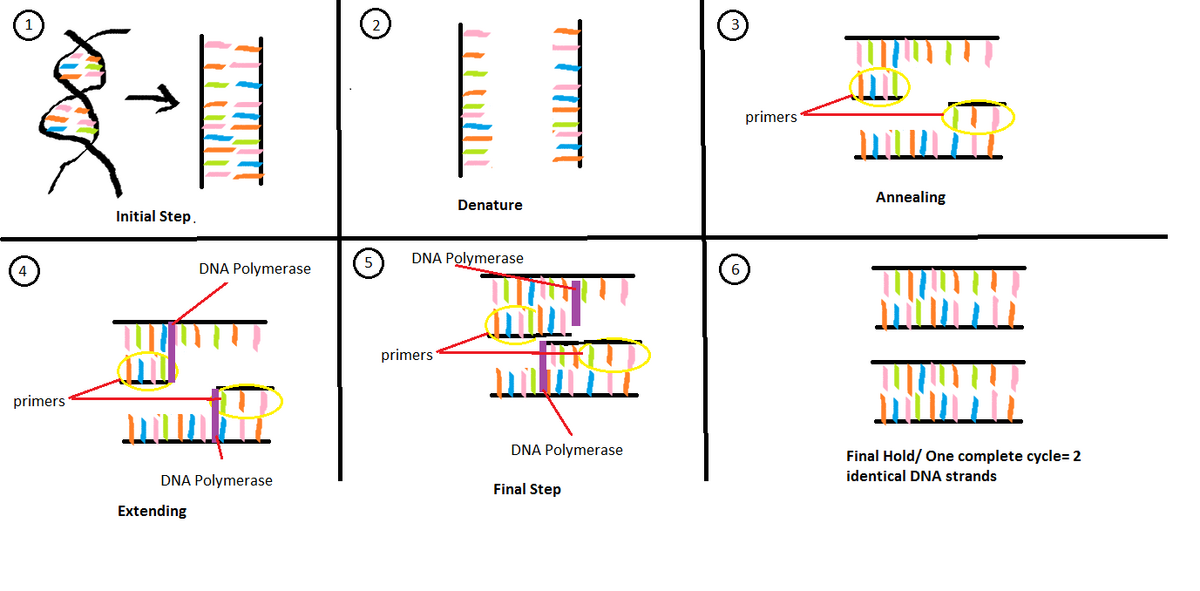
Components of a PCR reaction include a template DNA, primers, taq polymerase, magnesium chloride and deoxyribonucleotides. Template DNA is the "master copy" used to replicate and make copies of DNA. Primers are created to match any kind of specific nucleotide sequence a complementary DNA strand needs. They attach to either end of a DNA segment they want to copy. Taq polymerase binds to the DNA strand and primer. It then reads DNA code and attaches matching nucleotides to create DNA copies. These nucleotides called deoxyribonucleotides are the building blocks of the DNA molecules.They are made up of a deoxyribose sugar, a base and a phosphate group. Magnesium chloride acts as a cofactor of taq polymerase during the PCR process. PCR Thermal Cycling Process The process of PCR includes 6 different phases: the initial step (95° C for 3 minutes), denature (95° C for 30 seconds), anneal (57° C for 30 seconds), extend (72° C for 30 seconds), the final step (72° C for 3 minutes), and the final hold (4° C). The initial step consists of the machine heating up for the PCR process to begin and the helix of the DNA unwinding. After 3 minutes, the process begins to denature and the DNA splits into 2 single-stranded DNA molecules for 30 seconds. Once the DNA is denatured, the DNA template is annealed where primers bind onto the DNA template. After that, the process begins to extend and the DNA polymerase is activated and it locates a primer attached to a single DNA strand and then starts to add complementary nucleotides. The extension process consists of four types of nucleotides (adenine (A), thymine (T), cytosine (C) and guanine (G)) where A pairs with T and G pairs with C. These four nucleotides are paired together by hydrogen bonding. This process allows primers to stick to the template strand. 30 seconds after the extension process, the process approaches the final step where the process continues to add nucleotides to strands and then the process cools down to 4° C (final hold) to conclude one cycle. Another cycle is repeated except using two DNA strands as templates. The diagram below summarizes this process: References Kaiser, Gary. "MIcrobial Genetics." The Community College of Baltimore. The Community College of Baltimore, n.d. Web. 30 Oct. 2013. <http://faculty.ccbcmd.edu/courses/bio141/lecguide/unit6/genetics/DNA/DNA/dna.html>. "PCR Virtual Lab." Learn Genetics: Genetic Science Learning Center. The University of Utah, n.d. Web. 30 Oct. 2013. <http://learn.genetics.utah.edu/content/labs/pcr/>. "What is PCR?." Open PCR. Open PCR, n.d. Web. 30 Oct. 2013. <http://openpcr.org/what-is-pcr>. Greenwald, Ted, "DNA Sequencing For Fun And Profit: A Low-Cost Platform For Garage Biotech" Forbes.com, Web.
| |