Synthetic Biology (Spring2008): Computer Modelling Practicals

<html>
<body>
<script type="text/javascript">
var sc_project=3315875;
var sc_invisible=0;
var sc_partition=36;
var sc_security="779debd0";
</script>
<script type="text/javascript" src="http://www.statcounter.com/counter/counter_xhtml.js"></script><noscript>
</noscript>
</body>
</html>
Example: A --> B
Toy model for this example
- We are considering a very simple reaction network.
- It takes a compound 'A' to be converted into a compound 'B'.
- Following mass action laws, the kinetic law associated to the reaction is a first order reaction, with parameter k1.
- Model described : [math]\displaystyle{ A \xrightarrow{k_{1}} B }[/math]
- Kinetic Law for 'A': [math]\displaystyle{ \frac{d[A]}{dt} = -k_{1}[A] ; [A]_{t=0} = A_{0} > 0 }[/math]
- Kinetic Law for 'B': [math]\displaystyle{ \frac{d[B]}{dt} = k_{1}[A] ; [B]_{t=0} = 0 }[/math]
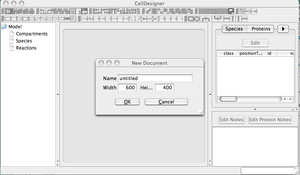
Starting a new CellDesigner Project
- Open CellDesigner Application: Double-click on the icon found on the desktop.
- Open a New Project: File --> New.
- Name your project 'test_1', for example. No need to change the dimensions of the work area.
|
|
Defining the reaction network
|
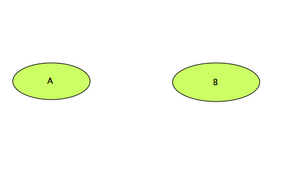
Main Working Area: Creating Compounds
- We want now to create 2 compounds: 'A' and 'B'.
- Click on the 'Simple Molecule' icon

- Click then in the working area, where you want to create the first compound. Name the compound 'A'.
- Repeat the previous operations to create compound 'B'.
- Save your project. File --> Save.
|
|
|
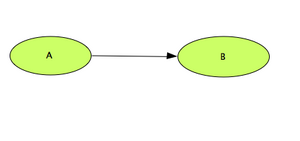
Main Working Area: Defining the reaction
- To create the reaction linking 'A' to 'B', click on the 'State Transition' icon

- Back to the working area, Click first on compound 'A', and then on compound 'B'. A reaction link is created.
- The topology of your reaction network is now created.
- Save your project. File --> Save.
|
|
Defining the reaction kinetic
|
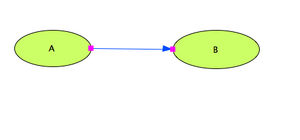
Main Working Area: Defining the kinetic law
- We want now to set the kinetic property of the defined reaction
- Right-click on the reaction link. It turns blue when you click on it.
- Select the 'Edit Reaction' fom the pop-up menu. A new window pops out with the reaction properties.
|
|
|
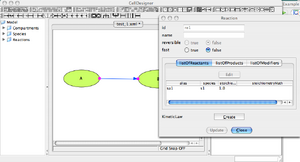
'Reaction' Window
- This window describes the attributes of the selected reaction link.
- A kinetic law has to be described to allow us to simulate the reactions.
- Click on: Kinetic Law 'Create'. A new window pops out.
|
|
|
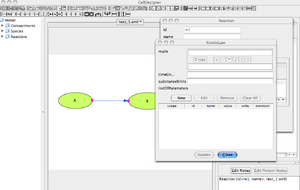
'Kinetic Law' Window
- The reaction link describes how reactants are converted into products.
- The kinetic law represents how the reactants reacts in time to form the products.
- First, you need to create a New parameter K1. Click on ListOfParameters NEW.
- A new window pops out.
|
|
|
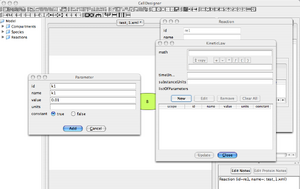
'Parameter' Window
- Set parameter id = k1, name = k1, initial value .01 . No units, Constant = True.
- Then click 'Add' to register the new parameter. And then, Click 'Close'.
- The window closes. You are now back to the 'Kinetic Law' Window.
|
|
|
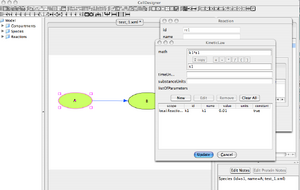
'Kinetic Law' Window
- The actual mathematical description of the reaction is now needed.
- Use the field called 'Math' to enter the kinetic law: [math]\displaystyle{ k_{1}[A] }[/math]
- [A] must be represented by its id, not its name !
- If you Click on the compound 'A' in the main working area, its 'id' will pop out in the Kinetic Law window.
- Once you are done, Click on 'Update'. And then 'Close'.
- Save your project. File --> Save.
|
|
Simulating the reactions
Main Simulation Features
|
Main Working Area
- Your reaction network is now ready to run a simulation.
- Select within the Menu 'Simulation' --> Control Panel
- The 'Control Panel' for the simulations pops out.
|
|
|
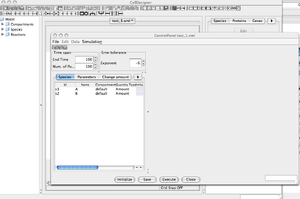
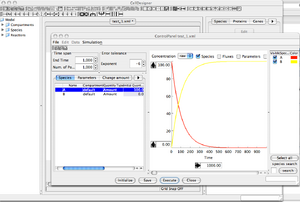
'Control Panel' Window
- At the moment, both compounds, 'A' and 'B', have a default initial value of zero. Nothing interesting could happen.
- Change 'A' initial amount from the 'Species' menu. Set it to 100.
- Set EndTime = 1000, NbPoints = 1000.
- You can now press the 'Execute' button to launch the simulation.
|
|
Interactive Simulation
|
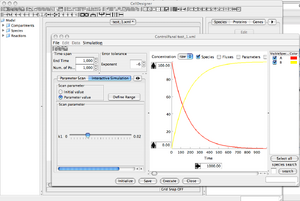
'Control Panel' Window
- This function helps you to vary a parameter, or a compound amount interactively.
- Select the 'Interactive Simulation' Menu, in the left area of the control panel.
- Select the 'parameter' button, and the slide menu gives you the option to vary k1.
- >>> more info
|
|
Parameter Scan
|
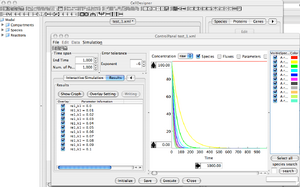
'Control Panel' Window
- Select the 'Parameter Scan' Menu
- It allows you to run multiple simulation while varying a specific parameter.
- This function is useful to run a simple sensitivity analysis on a given model.
- Check how the k1, or initial amount of A, are impacting the simulations.
- >>> more info
|
|
Additional features
To finish this first part of the tutorial, take the time to explore some features that will be useful for you to write your reports:
- Save image
- Export data
- >> more info from the CellDesigner Tutorial.












